Edepot.wur.nl
APPLIED AND ENVIRONMENTAL MICROBIOLOGY, Mar. 2008, p. 1394–1401
0099-2240/08/$08.00⫹0 doi:10.1128/AEM.01463-07Copyright 2008, American Society for Microbiology. All Rights Reserved.
Two Different Tetracycline Resistance Mechanisms, Plasmid-Carried
tet(L) and Chromosomally Located Transposon-Associated
tet(M),
Coexist in
Lactobacillus sakei Rits 9䌤
Mohammed Salim Ammor,1§ Miguel Gueimonde,1 Morten Danielsen,2 Monique Zagorec,3
Angela H. A. M. van Hoek,4 Clara G. de los Reyes-Gavila
Baltasar Mayo,1 and Abelardo Margolles1*
Instituto de Productos La
´cteos de Asturias (CSIC), Carretera de Infiesto s/n, 33300 Villaviciosa, Asturias, Spain1
; Chr. Hansen A/S,
Bøge Alle
´ 10-12, 2970 Hørsholm, Denmark2
; Unite
´ Flore Lactique et Environnement Carne
´, UR309, INRA, Domaine de Vilvert,
F-78350 Jouy-en-Josas, France3
; and RIKILT – Institute of Food Safety, Wageningen UR,
Bornsesteeg 45, Wageningen, The Netherlands4
Received 30 June 2007/Accepted 26 December 2007
Lactobacillus sakei is extensively used as functional starter culture in fermented meat products. One of the
safety criteria of a starter culture is the absence of potentially transferable antibiotic resistance determinants.
However, tetracycline-resistant L. sakei strains have already been observed. In this paper, we show that
tetracycline resistance in L. sakei Rits 9, a strain isolated from Italian Sola cheese made from raw milk, is
mediated by a transposon-associated tet(M) gene coding for a ribosomal protection protein and a plasmid-
carried tet(L) gene coding for a tetracycline efflux pump. pLS55, the 5-kb plasmid carrying the tet(L) gene, is
highly similar to the pMA67 plasmid recently described for Paenibacillus larvae, a species pathogenic to
honeybees. pLS55 could be transferred by electroporation into the laboratory strain L. sakei 23K. While the L.
sakei 23K transformant containing pLS55 displayed an intermediate tetracycline resistance level (MIC, <32
g/ml), L. sakei Rits 9, containing both tetracycline-resistant determinants, had a MIC of <256
g/ml,
suggesting that Tet L and Tet M confer different levels of resistance in L. sakei. Remarkably, in the absence of
tetracycline, a basal expression of both genes was detected for L. sakei Rits 9. In addition, subinhibitory
concentrations of tetracycline affected the expression patterns of tet(M) and tet(L) in different ways: the
expression of tet(M) was induced only at high tetracycline concentrations, whereas the expression of tet(L) was
up-regulated at lower concentrations. This is the first time that two different mechanisms conferring resistance
to tetracycline are characterized for the same strain of a lactic acid bacterium.
Lactobacillus sakei is a facultative heterofermentative psy-
transfer between industrial bacterial species and food-borne
chrotrophic lactic acid bacterium (LAB) that has been isolated
pathogens. Therefore, a consensus criterion has been issued
from several raw fermented food products of plant and animal
for which strains to be used in food systems should be free of
origin. It is found in kimchi, silage, cheese, sauerkraut, sour-
potentially transferable antibiotic resistance traits (15).
dough, and smoked fish but is mainly found in meat products
Tetracyclines are a group of broad-spectrum antibiotics
(4, 7, 8). Though some
L.
sakei strains have been identified as
whose general usefulness has been reduced with the onset of
responsible for the spoilage of vacuum-packaged meat prod-
bacterial resistance. Tetracycline resistance (Tcr) is the most
ucts, this bacterium is widely used as a starter culture for the
frequent bacterial antibiotic resistance found in nature and is
production of fermented sausages and has biotechnological
mostly acquired by horizontal gene transfer. Nowadays, 39
potential for biopreservation and food safety (6). Lactobacilli
acquired tetracycline determinants are known for bacteria
are generally recognized as safe and they are not responsible
(37). Usually, these genes code for energy-dependent efflux
for human infections in healthy people (46). However, they
systems or for proteins that protect the bacterial ribosomes
might act as reservoirs of transmissible antibiotic resistance
from the blockage of protein synthesis (9, 10, 37). In rare cases,
genes that under certain conditions could be transferred to
Tcr is mediated through direct inactivation of the antibiotic
food or gut microbiota (27). In addition, the emergence of
(40) or by mutations in the 16S rRNA that prevent the binding
antibiotic-resistant food-borne pathogens originating from
of tetracycline to the ribosome (38).
meat products (14) raises the question of the possibility of gene
Currently, data on antibiotic resistance in lactobacilli are
relatively scarce. However, in recent years a number of studieshave correlated atypically high phenotypic resistances with the
* Corresponding author. Mailing address: Instituto de Productos
presence of
tet genes (11, 17, 18, 19, 20, 26). Tetracycline
´cteos de Asturias, Consejo Superior de Investigaciones Cientı´ficas
resistance in
Lactobacillus has commonly been associated with
(IPLA-CSIC), Ctra. Infiesto s/n, 33300 Villaviciosa, Asturias, Spain.
Phone: 34 985 89 21 31. Fax: 34 985 89 22 33. E-mail: amargolles@ipla
the presence of
tet(M) (19, 20), but recently the gene coding
for the efflux transporter Tet L was also described for some
§ Present address: Laboratory of Microbiology and Biotechnology of
cloacal isolates (5). However, data about the functionality of
Foods, Agricultural University of Athens, 75 Iera Odos, 11855 Athens,
both genes when they coexist in the same bacterium were not
䌤 Published ahead of print on 11 January 2008.
available until now. In this context, this study reports the iso-
TETRACYCLINE RESISTANCE IN
L.
SAKEI Rits 9
TABLE 1. Primers used in this study
Sequence (5⬘–3⬘)
Reference or source
J. M. Collard, personal
tet(M) flanking regions
tet(M) flanking regions
a R, reverse; F, forward.
b ITS, internal transcribed spacer.
lation of a Tcr
L.
sakei strain from Italian Sola cheese and the
(33). Total RNA was extracted from cells grown up to an OD
molecular characterization of both ribosomal protection- and
use of an RNeasy mini kit (Qiagen) following the manufacturer's instructionswith the following modifications: the lysis buffer was supplemented with 30 g/ml
efflux pump-encoding genes,
tet(M) and
tet(L), responsible for
lysozyme (Sigma) and 100 U/ml mutanolysin (Sigma) and the samples were
Tcr in this strain.
incubated for 30 min under gentle stirring. DNA was removed by on-columndigestion using an RNase-free DNase set (Qiagen). Four microliters of RNA
MATERIALS AND METHODS
(about 3 g) was reverse transcribed into cDNA by use of a cDNA archive kit(Applied Biosystems, Foster City, CA). The cDNA was stored at ⫺80°C until
Bacterial strains and growth conditions. L.
sakei Rits 9 was isolated on MRS
agar (Oxoid Limited, Hampshire, United Kingdom) containing 16 g/ml tetra-
(ii) Microarray hybridization. DNA microarrays contained 327 oligonucleo-
cycline (Sigma, St. Louis, MO) from an Italian Sola cheese made from raw cow's
tides (50 to 60 base pairs long), including control probes and oligonucleotides
milk according to International Dairy Federation (IDF) standard 122C:1996. An
specific for 250 antibiotic resistance genes, including 28
tet genes (1). Spotting of
internal 474-bp fragment of the 16S rRNA gene and an internal 424-bp fragment
the oligonucleotides, hybridization conditions, and analysis of the results were as
of the
katA gene (encoding the
L.
sakei heme-dependent catalase) were ampli-
previously described (43).
fied using primers Y1 (45) and R518 and primers 702-F and 310-R (2), respec-
(iii) Real-time PCR conditions. Real-time PCR was used to assess the influ-
tively (Table 1). The resulting nucleotide sequences showed to be identical to the
ence of different subinhibitory concentrations of tetracycline (16, 32, and 64
corresponding partial sequences in
L.
sakei 23K (6).
L.
sakei 23K, a laboratory
g/ml) on the expression levels of
tet(L) and
tet(M) in
L.
sakei Rits 9. All the
strain originally isolated from sausage and cured of plasmids (3), was used as the
primers used in this study are listed in Table 1. Primers TetL-FW-RT and
recipient strain for genetic constructions.
L.
sakei 23K electrocompetent cells
TetL-RV-RT and TetM-FW-RT and TetM-RV-RT were designed to amplify
were prepared and transformed with pLS55 as described previously (3). After anincubation period of 2 h following electroporation, bacterial suspensions were
internal fragments of 70 and 78 bp, respectively. The rRNA 16S-to-23S inter-
plated on MRS medium containing 4, 8, 16, or 32 mg/liter tetracycline and
genic region was used as the endogenous control by using
Lactobacillus-specific
incubated for 48 h at 30°C.
primers (24). PCR was performed in an ABI Prism 7500 fast real-time PCR
Bacterial strains were stored at ⫺80°C and routinely cultured on MRS agar.
system (Applied Biosystems), and SYBR green I fluorophore was used to cor-
All incubations were performed aerobically at 30°C for 48 h.
relate the amount of PCR product with the fluorescent signal. Amplification was
Determination of the MICs of tetracycline. The MICs of tetracycline for the
carried out in a 25-l final volume containing 1 l of cDNA as a template, 200
different strains were determined by microdilution. Briefly, colonies obtained
nM of each primer, and 12.5 l of SYBR green PCR master mix (Applied
after growth on solid media were picked up and incubated overnight at 30°C in
Biosystems). Thermal cycling consisted of an initial cycle of 95°C for 10 min
LSM broth (29). The optical density at 625 nm (OD
) of the cultures was
followed by 35 cycles of 95°C for 15 s and 60°C for 1 min. The expression levels
adjusted to 0.2 in LSM broth, and the suspension was diluted 500-fold in the
in the presence of antibiotic were refereed to those obtained for the control
same medium. One hundred microliters of this dilution was then transferred to
culture (absence of antibiotic). Two independent experiments were carried out
100 l of LSM containing the appropriate amount of tetracycline in serial
and each sample was analyzed in duplicate in two independent PCR runs.
twofold dilutions, and the microtiter plates were incubated at 30°C for 24 h. The
Negative controls, including all the elements of the reaction mixture except the
growth was recorded with a Benchmark plus microplate spectrophotometer
template cDNA, were also included.
(Bio-Rad, Hercules, CA). All the experiments were carried out in triplicate.
(iv) Pulsed-field gel electrophoresis (PFGE) and Southern hybridization con-
DNA and RNA techniques. (i) Nucleic acids extractions and labeling.
ditions. The genetic location of
tet(L) and
tet(M) was assessed by hybridization
Genomic DNA was isolated using the GenElute bacterial genomic DNA kit
using as probes 0.7- and 1.5-kb internal segments of the genes obtained by PCR
(Sigma). Plasmid DNA was isolated using either the large-scale Qiagen kit
and labeled with digoxigenin (Roche Applied Science, Basel, Switzerland). The
(Qiagen Inc. Valencia, CA) or the procedure of O'Sullivan and Klaenhammer
tet(L) and
tet(M) fragments were amplified using primer pairs TetL-FW3/TetL-
AMMOR ET AL.
APPL. ENVIRON. MICROBIOL.
RV3 and DI/TetM-R (9), respectively. Total and plasmid DNAs digested withthe restriction enzymes EcoRI, HindIII, AscI, and PstI (Takara Bio Inc., Shiga,Japan) were hybridized using high-stringency standard conditions at 68°C.
For PFGE analysis, the strain was inoculated in 10 ml MRS supplemented with
20 mM DL-threonine and incubated at 30°C until the OD
was 0.5 to 1.0 or
above. The cells were harvested by centrifugation, washed in 10 ml 50 mMEDTA, and resuspended in 50 mM EDTA (300 l ⫻ OD
). A 125-l cell
suspension was mixed gently with 750 l 1% low-melting-point agarose (pre-pared in 50 mM EDTA). The cell-agarose suspension was pipetted into theBio-Rad plug mold. The agarose plugs were incubated at 37°C overnight in alysozyme solution (2 mg/ml lysozyme, 20 units/ml mutanolysin, 0.05% N-lauroylsarcosine in 50 mM EDTA). The lysozyme solution was replaced by a sodiumdodecyl sulfate-proteinase solution (10 mM Tris, pH 8.0, 1% sodium dodecylsulfate, 2 mg/ml proteinase K in 0.5 M EDTA, pH 8.5) and incubated at 50°Covernight. The agarose plugs were washed six times for 30 min in 50 mM EDTAand stored at 4°C in 50 mM EDTA. Slices of 1 to 2 mm of the agarose plugs wereincubated in 200 l of restriction enzyme buffer for 1 to 4 h at 4°C. The buffer wasreplaced with 200 l fresh restriction enzyme buffer, 2 l acetylated bovineserum albumin (10 mg/ml stock), and 20 to 40 units of AscI. The agarose plugswere incubated for 30 to 45 min at 4°C and then at 37°C overnight. The sampleswere loaded on a 1.1% agarose gel prepared in 0.5⫻ Tris-borate-EDTA buffer.
The DNA fragments were resolved on a Bio-Rad contour-clamped homoge-neous electric field mapper using a 24-h program with a linear ramp factor, aninitial switch time of 2 s, and a final switch time of 30 s. The gel was stained inethidium bromide and destained in 0.5⫻ Tris-borate-EDTA buffer.
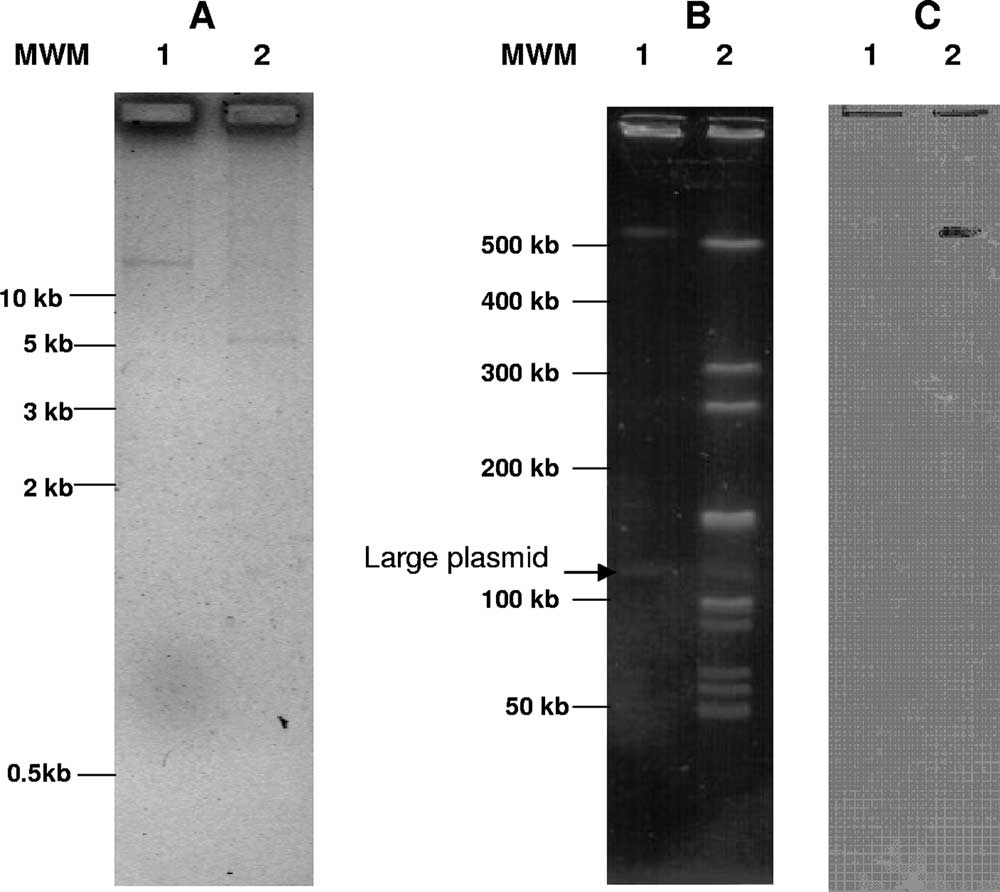
FIG. 1. (A) Plasmid profile of L. sakei Rits 9 undigested (lane 1)
Southern blotting of PFGE gels was performed with DNA probes labeled with
and digested with PstI (lane 2). (B) PFGE analysis of total DNA from
horseradish peroxidase with the ECL direct nucleic acid labeling kit (Amersham
L. sakei Rits 9 undigested (lane 1) and digested with AscI (lane 2).
Biosciences, Buckinghamshire, United Kingdom) according to the manufactur-
(C) Southern blot analysis of the PFGE gel with the internal
er's instructions.
tet(M) probe. MWM, molecular weight marker.
(v) Sequencing strategy for the tet genes and sequence analysis. Plasmid DNA
was sequenced after serial runs using the first-round primers, which consisted ofthe complementary sequences of TetL-FW3 and TetL-RV3 and then primersdesigned from the DNA sequence newly obtained. The plasmid was thereafterresequenced on the other strand in order to check for sequence accuracy.
signal (data not shown). To verify the presence of both genes,
For sequencing the tet(M) region, a pair of primers was designed from the
primers derived from known tet gene sequences were used.
tet(M) sequence of Staphylococcus aureus subsp. aureus Mu50 and served for the
Amplification of internal fragments of tet(L) and tet(M) with
amplification of L. sakei Rits 9 tet(M). Primers tetM-revF and tetM-revR (Table1) were used to amplify regions upstream and downstream of the tet(M) genes.
the primers TetL-FW3 and TetL-RV3 and DI and TetM-R,
The sequencing of the flanking regions of tet(M) was carried out using inverse
respectively, resulted in amplicons of about 0.7 kb and 1.5 kb,
PCR as described elsewhere (16). In short, total genomic DNA was digested with
confirming that L. sakei Rits 9 possesses both genes. L. sakei
HindIII and self-ligated overnight. The ligated DNA was precipitated, centri-
Rits 9 harbors one small plasmid of 5 kb, as revealed by a
fuged, dried, and resuspended in 100 l Tris-EDTA prior to use as the template
plasmid profile analysis using the O'Sullivan and Klaenham-
for PCR amplification. Purified PCR products were sequenced by cycle extensionin an ABI 370 DNA sequencer (Applied Biosystems).
mer method (Fig. 1A), and at least one large plasmid, as
Phylogenetic analyses were performed on sequences available in the GenBank
revealed by PFGE (Fig. 1B). Southern blots showed tet(L) to
database, using the Treetop software (http://www.genebee.msu.su/services
be located on the 5-kb small plasmid (data not shown) and
tet(M) on a large AscI PFGE chromosomal fragment (⬎450
Nucleotide sequence accession numbers. The nucleotide sequences described
in this paper have been deposited in the GenBank database with the following
kb) (Fig. 1C). In order to determine the involvement of those
accession numbers: for L. sakei Rits 9 plasmid pLS55, EF605268; and for L. sakei
two genes in the resistance phenotype of L. sakei Rits 9, the
Rits 9 tet(M) and flanking regions, EF605269.
5-kb plasmid containing tet(L) was totally sequenced, as wasthe chromosomal region encompassing tet(M).
The tet(L) gene is contained by a plasmid, and the tet(M)
gene is flanked by transposon-like regions. The 5-kb plasmid
L. sakei Rits 9 possesses tet(L) and tet(M) resistance genes.
containing tet(L), named pLS55, was sequenced. It was found
L. sakei Rits 9 was isolated from an Italian Sola cheese as
to be composed of 5,031 bp, consistent with its predicted size.
spontaneously resistant to tetracycline. The presence of genes
The plasmid was almost 100% identical to pMA67, a plasmid
responsible for such resistance was searched by hybridization
recently described for the gram-positive bacterial pathogen of
with DNA microarrays containing oligonucleotides character-
honeybees Paenibacillus larvae (32). Indeed, only seven of the
istic of 28 known tetracycline resistance genes. The results
base pairs were found to be different, four of them located in
showed the strain to harbor both tet(M) and tet(L). Hybridiza-
the tet(L) structural gene (positions 1, 287, 859, and 1197), and
tion signals were quite strong for both 50- and 60-mer oligo-
the plasmids differ in size by only one nucleotide (5,030 bp for
nucleotides used for identifying the respective Tcr genes. Ex-
pMA67). Remarkably, a different initiation codon was found
cept for positive signals obtained with control probes targeting
for tet(L) in pLS55 (ATG instead of GTG), which could sug-
lactobacillus tuf genes, no other positive signals were found
gest a more efficient translation of the gene in L. sakei (30).
with any of the remaining spots, indicating the absence of other
The expression of tet(L) seems to depend on the synthesis of a
antibiotic resistance determinants. This shows either that other
20-amino-acid leader peptide encoded 22 bp upstream of the
resistance genes are absent or that similar genes may be
tet(L) ribosome binding site, which is typical of inducible tet
present but with a homology too low to get a hybridization
genes (25). A phylogenetic analysis performed on all complete
TETRACYCLINE RESISTANCE IN L. SAKEI Rits 9
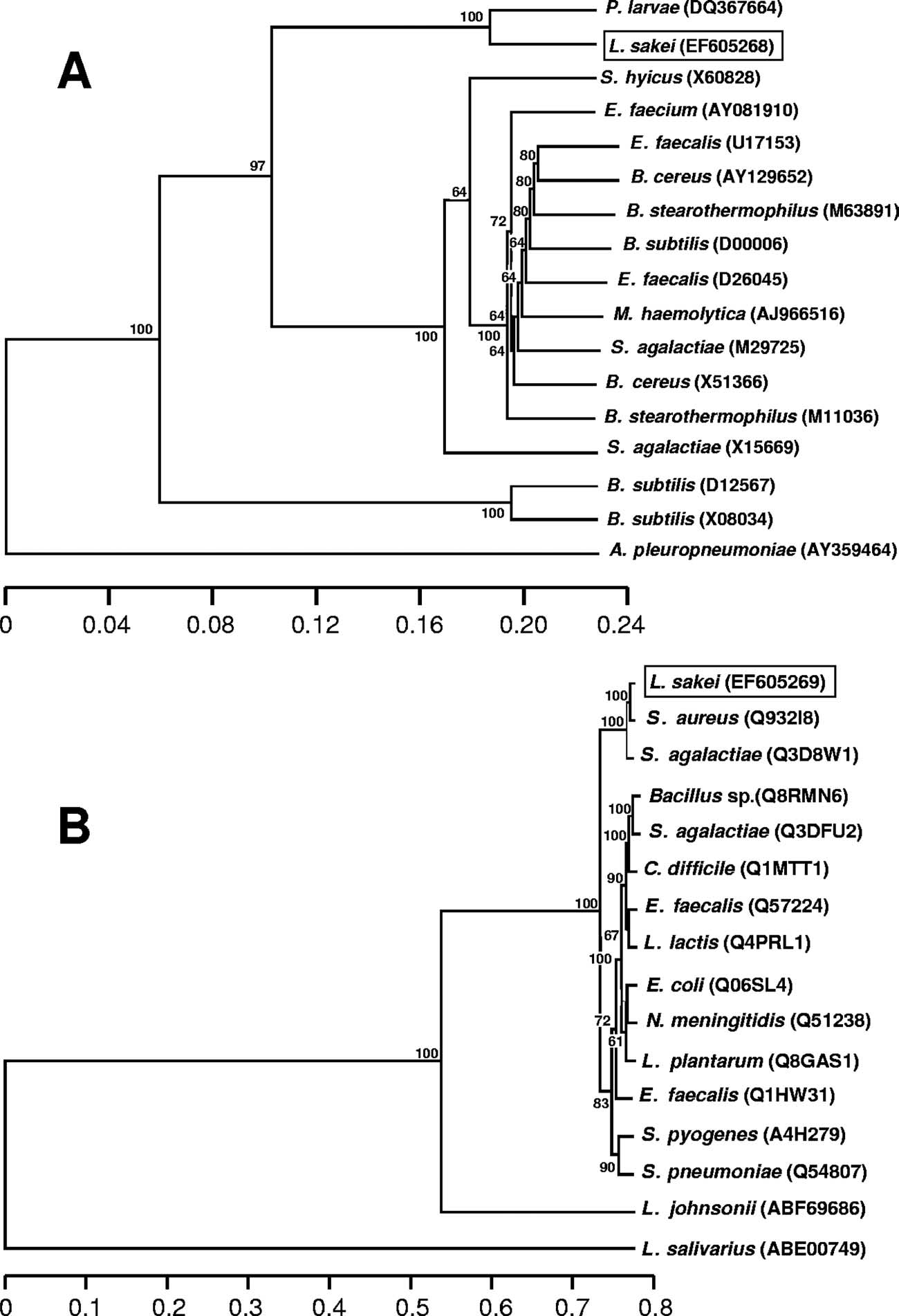
FIG. 2. Phylogenetic tree of homologs of the deduced Tet(L) and Tet(M) proteins (A and B, respectively). Protein accession numbers are given
in brackets. Trees were constructed by the neighbor-joining algorithm and clustered by the unweighted-pair group method using average linkages,and bootstrap values (100 replicates) are given at the branch points. The distances refer to the percentages of different residues. Abbreviations:E. faecium, Enterococcus faecium; B. cereus, Bacillus cereus; M. haemolytica, Mannheimia haemolytica; A. pleuropneumoniae, Actinobacilluspleuropneumoniae; C. difficile, Clostridium difficile; E. coli, Escherichia coli; N. meningitidis, Neisseria meningitidis.
tet(L) sequences available in the GenBank database showed
deduced that pLS55 would likely be a mobilizable rolling-circle
that both P. larvae and L. sakei Rits 9 tet(L) genes are different
replication plasmid in the group II family (also called the
from all previously described tet(L) genes and form an inde-
pMV158 family).
pendent branch associated with a very strong bootstrap value
The sequence of a region encompassing 8,524 bp around the
tet(M) gene was obtained by several PCR and sequencing
Apart from tet(L), pLS55 contains all the elements for rep-
steps. The nucleotide sequence of the L. sakei Rits 9 tet(M)
lication control (12, 13, 22, 28) (Fig. 3A). Interestingly, the Rep
gene was shown to be identical to the one described for S.
protein is 80% identical to the Rep proteins of L. sakei plasmid
aureus subsp. aureus Mu50 and for Streptococcus agalactiae
pLS141-1 and of pLC2 identified for Lactobacillus curvatus, a
COH1 (Fig. 2B). The tet(M) gene was flanked downstream and
lactobacillus species closely related to L. sakei (GenBank ac-
upstream by regions with high similarity to the tet(M)-sur-
cession no. AB109041 and CAA78602, respectively). It can be
rounding regions of several gram-positive bacteria (31, 36, 41),
AMMOR ET AL.
APPL. ENVIRON. MICROBIOL.
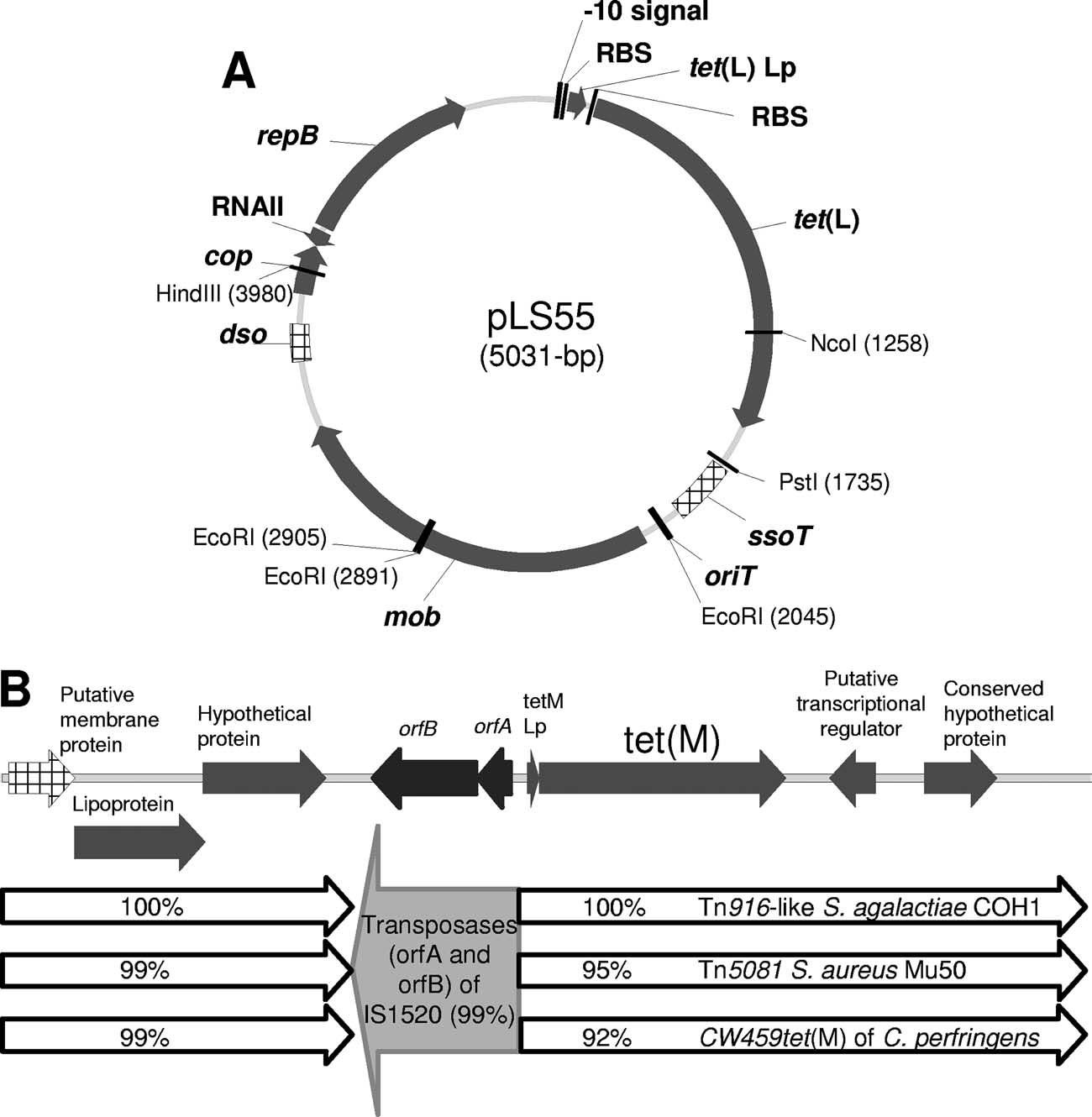
FIG. 3. Genetic structure of the tet(L)-containing plasmid pLS55 (A) and L. sakei Rits 9 tet(M) and flanking regions (B). Arrows show the
direction of transcription of the open reading frames. Relevant restriction sites and their locations are indicated. The genes which matched thehighest homology scores and the homologies with the partial sequences of different transposons are indicated.
corresponding to transposon-like sequences (Fig. 3B). Up-
higher tetracycline concentrations (between 2.74 ⫾ 0.40- and
stream of tet(M), a 181-bp region mainly features a sequence
3.07 ⫾ 0.44-fold increases) (Fig. 4).
corresponding to a 28-amino-acid leader peptide. Immediately
pLS55 is able to replicate in L. sakei 23K. To determine
upstream the leader peptide sequence, we found a 1,305-bp
whether pLS55 replication is possible in another L. sakei strain,
sequence that shares more than 99% identity with L. sakeiIS1520, encompassing the transposase subunits A and B of anIS element present in five copies in the L. sakei 23K chromo-some (6).
Regulation and expression levels of tet(M) and tet(L). Real-
time PCR was used to assess the influence of different subin-hibitory concentrations of tetracycline (16, 32, and 64 g/ml)on the expression levels of tet(M) and tet(L) in L. sakei Rits 9.
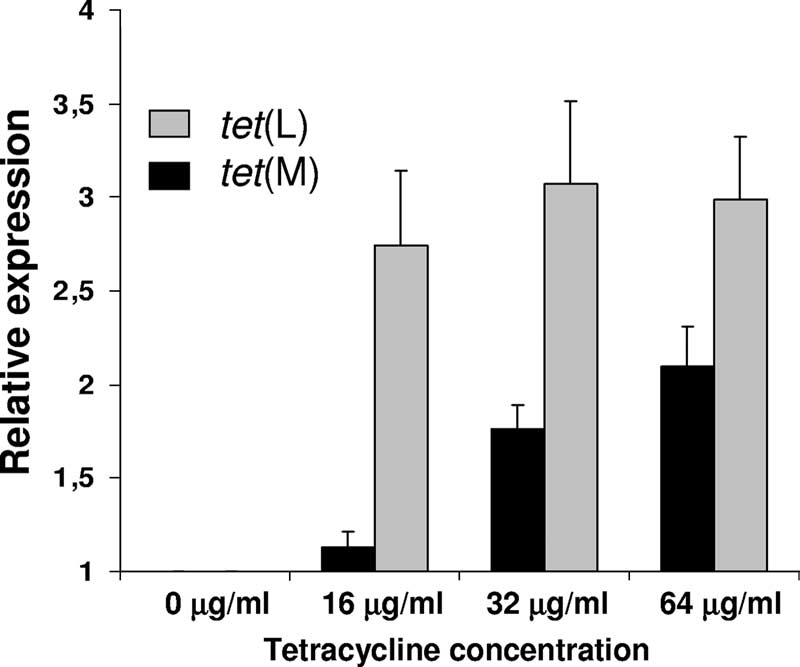
Concentrations higher than 64 g/ml affected the growth rateof the strain and therefore were not included in the study. Abasal constitutive expression of both genes was observed inde-pendent of the presence of tetracycline. Remarkably, we no-ticed that tet(M) expression was gradually induced by exposureto increasing amounts of tetracycline. Indeed, tet(M) inductionwas about 13% increased at low tetracycline concentration (16g/ml) and up to 100% (relative induction was 2.095 ⫾ 0.215)after exposure to 64 g/ml compared to the control conditions
FIG. 4. Relative expression levels of tet(L) and tet(M) in L. sakei
(absence of antibiotic) (Fig. 4). On the contrary, the tet(L)
Rits 9 grown in the presence of different tetracycline concentrations
gene was induced up to 2.74-fold at the lower tetracycline
refereed to those obtained for the control culture (absence of anti-
concentration, and its relative expression remained similar at
TETRACYCLINE RESISTANCE IN L. SAKEI Rits 9
and to assess the functionality of the tet(L) gene, the transfor-
pLS55 and the 80% identity between the Rep protein of pLS55
mation of the plasmid into L. sakei 23K was attempted, and
and some Rep proteins described for other L. sakei or L.
transformants were plated with different tetracycline concen-
curvatus plasmids suggest that pLS55 can be transferred and
trations (4, 8, 16, and 32 g/ml). When 4 g/ml was used, a
stably maintained in L. sakei. Indeed, we could electroporate it
background was quite visible, but the background disappeared
in the plasmid-free laboratory strain L. sakei 23K, in which it
when 8-, 16-, and 32-g/ml concentrations of tetracycline were
used. Several transformants were obtained on plates with 8 and
On the other hand, tet(M) was shown to be located on a
16 g/ml of tetracycline. No transformants were obtained at 32
transposon-like region. Upstream of tet(M), a fragment of
g/ml. Plasmid preparations of four clones confirmed the pres-
1,305 bp identical to L. sakei IS1520 was also present. This
ence of a 5-kb plasmid in all of them. Then, one of them,
suggests that the acquisition of tet(M) by L. sakei Rits 9 oc-
named L. sakei 23K-TL, was selected to analyze its MIC to
curred through an insertion event, although a more detailed
tetracycline in comparison with the control L. sakei 23K and L.
study is necessary to corroborate this.
sakei Rits 9. While the MIC of the Rits 9 strain was found to
The high Tcr level in L. sakei Rits 9 and the absence of
be ⬍256 g/ml and that of the 23K strain ⬍1 g/ml of tetra-
positive hybridization results other than the ones obtained with
cycline, the MIC of L. sakei 23K-TL was ⬍32 g/ml.
the tet(L) and tet(M) oligonucleotides in the microarray anal-ysis suggest that Tcr in this strain is linked to the presence of
one or both genes. In order to ascertain the functionalities ofboth genes and the partial contribution of each to the Tcr
Tetracyclines have been extensively used in the prophylaxis
phenotype, we have transformed the plasmid-free laboratory
and treatment of human and animal infections. Furthermore,
strain L. sakei 23K with the tet(L)-containing plasmid pLS55.
they have been administered at subtherapeutic concentrations
The resulting strain displayed an intermediate Tcr level com-
as growth promoters in animal feeds (34, 44). This intensive
pared with the Rits 9 strain, which displayed a much higher
and extensive use has caused Tcr to spread to a large number
MIC. Thus, the higher Tcr level of L. sakei Rits 9 could be due
of commensal bacteria (9, 37). In fact, different Tcr genes are
to the presence of tet(M) or to a synergistic effect of both
present in the fecal microbiota of babies not previously ex-
genes. Furthermore, these data indicate that both tet genes are
posed to the antibiotic (23). At present, there is great concern
functional in L. sakei, with tet(L) conferring a moderated re-
that animal and human commensal bacteria, such as LAB,
sistance level, whereas tet(M) confers a high Tcr level to this
could act as a reservoir for antibiotic resistance genes. These
bacteria. In relation to this, it has been shown that tet(L) and
microorganisms may subsequently contaminate the raw milk
tet(M) can contribute differently to the Tcr phenotype depend-
and meat produced from these animals, and the foods pre-
ing on the Enterococcus or Streptococcus strain (39). It is also
pared from those raw materials can therefore be considered as
likely that the resistance level conferred by these two genes is
potential vehicles for the spread of antibiotic-resistant LAB
species dependent and probably strain dependent.
along the food chain to the consumer (42). Resistances could
Finally, expression studies were carried out to go more
ultimately be transferred to human pathogenic and opportu-
deeply into the functionality of tet(L) and tet(M) in L. sakei
nistic bacteria, hampering the treatment of infections (27).
Rits 9. The fact that tet(M) expression was mainly induced at
Several Tcr LAB have been isolated from raw milk dairy
high Tcr levels, whereas tet(L) induction was achieved at lower
products, e.g., Lactobacillus fermentum ROT1 (21) and Lacto-coccus lactis subsp. lactis K214 (34), and from raw meat-based
concentrations, sheds some light onto the physiological func-
fermented products, such as L. alimentarius, L. curvatus, L.
tion of both genes. These data indicate that, at a low tetracy-
plantarum, and L. sakei (19). The Tcr has been found to be
cline concentration, the activity of the efflux pump Tet L is
mediated mainly by tet(M), which could be plasmid encoded
enough for L. sakei Rits 9 to cope with antibiotic challenge;
and transferred through interspecies and intergenus conjuga-
however, at concentrations higher than 16 g/ml, the cells need
tion mechanisms (17, 27). In this study, we show that L. sakei
an extra input, which is supplied by a higher amount of the
Rits 9, a Tcr strain isolated from a dairy product, harbors two
ribosomal protection protein Tet M. These findings also sup-
Tcr genes, namely, the ribosomal protection tet(M) gene fre-
port the previous results just discussed above, indicating that
quently encountered in lactobacilli and the efflux pump-encod-
Tet M is responsible, to a larger extent than Tet L, for the high
ing tet(L) gene. This combination of tet(L) and tet(M) genes is
Tcr phenotype of L. sakei Rits 9.
very frequently found for Streptococcus spp. and Enterococcus
In conclusion, the results of the current study indicate that
sp. strains (35, 39) and also for cloacal Lactobacillus salivarius
Lactobacillus species from raw milk cheese can harbor ac-
subsp. salivarius isolates (5). However, to the best of our
quired Tcr determinants associated with mobile elements, po-
knowledge this is the first report on the coexistence of two
tentially enabling them to spread to other LAB or potentially
genes encoding different mechanisms of Tcr in the same L.
pathogenic bacteria. We also demonstrated, for the first time,
that two different Tcr mechanisms, active efflux and ribosomal
The gene tet(L) was found to be associated with the plasmid
protection, are functional when they are together in the same
pLS55, which is highly similar to pMA67, a plasmid described
strain. Remarkably, our data suggest that the two genes are
for the honeybee-pathogenic species P. larvae (32). As L. sakei
dedicated to cope with two different physiological conditions,
and P. larvae are not known to share a common ecological
low and high tetracycline concentrations. This functional
niche, it is therefore plausible that such a plasmid has been
complementarity of both mechanisms and their involvement in
horizontally transferred in these two hosts through different
the physiology of L. sakei under tetracycline challenge will
microorganisms. The presence of a Mob protein encoded by
contribute to an understanding of how a bacterium makes use
AMMOR ET AL.
APPL. ENVIRON. MICROBIOL.
of different resistance determinants and of how they are en-
tet genes along the process line of fermented dry sausages. Syst. Appl.
gaged to fight against the deleterious action of antimicrobials.
20. Gevers, D., M. Danielsen, G. Huys, and J. Swings. 2003. Molecular charac-
terization of tet(M) genes in Lactobacillus isolates from different types of
fermented dry sausage. Appl. Environ. Microbiol. 69:1270–1275.
21. Gfeller, K. Y., M. Roth, L. Meile, and M. Teuber. 2003. Sequence and genetic
Work on antibiotic resistance at our laboratories was supported by
organization of the 19.3-kb erythromycin- and dalfopristin-resistance plas-
an EU project within the Sixth Framework Programme (ACE-ART,
mid pLME300 from Lactobacillus fermentum ROT1. Plasmid 50:190–201.
reference no. FP6-506214). M. S. Ammor was awarded a postdoctoral
22. Grohmann, E., G. Muth, and M. Espinosa. 2003. Conjugative plasmid trans-
fellowship from the Secretarı´a de Estado de Universidades e Investi-
fer in gram-positive bacteria. Microbiol. Mol. Biol. Rev. 67:277–301.
´n of the Spanish Ministry of Education and Science (reference
23. Gueimonde, M., S. Salminen, and E. Isolauri. 2006. Presence of specific
antibiotic (tet) resistance genes in infant faecal microbiota. FEMS Immunol.
no. SB2004-0165). Miguel Gueimonde is the recipient of a Juan de
Med. Microbiol. 48:21–25.
la Cierva contract from the Spanish Ministry of Education and
24. Haarman, M., and J. Knol. 2006. Quantitative real-time PCR analysis of
fecal Lactobacillus species in infants receiving a prebiotic infant formula.
Appl. Environ. Microbiol. 72:2359–2365.
25. Hoshino, T., T. Ikeda, N. Tomizuka, and K. Furukawa. 1985. Nucleotide
1. Ammor, M. S., A. B. Flo
´rez, A. H. A. M. van Hoek, C. G. de los Reyes-
sequence of the tetracycline resistance gene of pTHT15, a thermophilic
´n, H. J. M. Aarts, A. Margolles, and B. Mayo. 2008. Molecular char-
Bacillus plasmid: comparison with staphylococcal TcR controls. Gene 37:
acterization of intrinsic and acquired antibiotic resistance in lactic acid bac-
teria and bifidobacteria. J. Mol. Microbiol. Biotechnol. 14:6–15.
26. Huys, G., K. D'Haene, and J. Swings. 2006. Genetic basis of tetracycline and
2. Ammor, M. S., E. Dufour, M. Zagorec, S. Chaillou, and I. Chevallier. 2005.
minocycline resistance in potentially probiotic Lactobacillus plantarum strain
Characterization and selection of Lactobacillus sake [sic] strains isolated
CCUG 43738. Antimicrob. Agents Chemother. 50:1550–1551.
from traditional dry sausage for their potential use as starter cultures. Food
27. Jacobsen, L., A. Wilcks, K. Hammer, G. Huys, D. Gevers, and S. R.
Andersen. 2007. Horizontal transfer of tet(M) and erm(B) resistance plas-
3. Berthier, F., M. Zagorec, M. Champomier-Verge
s, S. D. Ehrlich, and F.
mids from food strains of Lactobacillus plantarum to Enterococcus faecalis
Morel-Deville. 1996. Efficient transformation of Lactobacillus sakei by elec-
JH2-2 in the gastrointestinal tract of gnotobiotic rats. FEMS Microbiol. Ecol.
4. Cappello, M. S., B. Laddomada, P. Poltronieri, and G. Zacheo. 2001. Char-
28. Khan, S. A. 1997. Rolling-circle replication of bacterial plasmids. Microbiol.
acterisation of lab in typical Salento Pecorino cheese. Meded. Rijksuniv.
Mol. Biol. Rev. 61:442–455.
Gent Fak. Landbouwkd. Toegep. Biol. Wet. 66:569–572.
29. Klare, I., C. Konstabel, S. Mu
¨ller-Bertling, R. Reissbrodt, G. Huys, M.
5. Cauwerts, K., F. Pasmans, L. A. Devriese, F. Haesebrouck, and A. Decostere.
Vancanneyt, J. Swings, H. Goossens, and W. Witte. 2005. Evaluation of new
2006. Cloacal Lactobacillus isolates from broilers often display resistance
broth media for microdilution antibiotic susceptibility testing of lactobacilli,
toward tetracycline antibiotics. Microb. Drug Resist. 12:284–288.
pediococci, lactococci, and bifidobacteria. Appl. Environ. Microbiol. 71:
6. Chaillou, S., M. C. Champomier-Verges, M. Cornet, A. M. Crutz-Le Coq,
A. M. Dudez, V. Martin, S. Beaufils, E. Darbon-Rongere, R. Bossy, V. Loux,
30. Kozak, M. 2005. Regulation of translation via mRNA structure in pro-
and M. Zagorec. 2005. The complete genome sequence of the meat-borne
karyotes and eukaryotes. Gene 361:13–37.
lactic acid bacterium Lactobacillus sakei 23K. Nat. Biotechnol. 23:1527–1533.
31. Kuroda, M., T. Ohta, I. Uchiyama, T. Baba, H. Yuzawa, I. Kobayashi, L. Cui,
s, M. C., S. Chaillou, M. Cornet, and M. Zagorec. 2002.
A. Oguchi, K. Aoki, Y. Nagai, J. Lian, T. Ito, M. Kanamori, H. Matsumaru,
Lactobacillus sakei: recent developments and future prospects. Res. Micro-
A. Maruyama, H. Murakami, A. Hosoyama, Y. Mizutani-Ui, N. K. Taka-
hashi, T. Sawano, R. Inoue, C. Kaito, K. Sekimizu, H. Hirakawa, S. Kuhara,
8. Choi, I. K., S. H. Jung, B. J. Kim, S. Y. Park, J. Kim, and H. U. Han. 2003.
S. Goto, J. Yabuzaki, M. Kanehisa, A. Yamashita, K. Oshima, K. Furuya, C.
Novel Leuconostoc citreum starter culture system for the fermentation of
Yoshino, T. Shiba, M. Hattori, N. Ogasawara, H. Hayashi, and K. Hira-
kimchi, a fermented cabbage product. Antonie van Leeuwenhoek 84:247–
matsu. 2001. Whole genome sequencing of meticillin-resistant Staphylococ-
cus aureus. Lancet 357:1225–1240.
9. Chopra, I., and M. Roberts. 2001. Tetracycline antibiotics: mode of action,
32. Murray, K. D., K. A. Aronstein, and J. H. de Leo
´n. 2007. Analysis of pMA67,
applications, molecular biology, and epidemiology of bacterial resistance.
a predicted rolling-circle replicating, mobilizable, tetracycline-resistance
Microbiol. Mol. Biol. Rev. 65:232–260.
plasmid from the honey bee pathogen, Paenibacillus larvae. Plasmid 58:89–
10. Clermont, D., O. Chesneau, G. De Cespedes, and T. Horaud. 1997. New
tetracycline resistance determinants coding for ribosomal protection in
33. O'Sullivan, D. J., and T. R. Klaenhammer. 1993. Rapid mini-prep isolation
streptococci and nucleotide sequence of tet(T) isolated from Streptococcus
of high-quality plasmid DNA from Lactococcus and Lactobacillus spp. Appl.
pyogenes A498. Antimicrob. Agents Chemother. 41:112–116.
Environ. Microbiol. 59:2730–2733.
11. Danielsen, M. 2002. Characterization of the tetracycline resistance plasmid
34. Perreten, V., F. Schwarz, L. Cresta, M. Boeglin, G. Dasen, and M. Teuber.
pMD5057 from Lactobacillus plantarum 5057 reveals a composite structure.
1997. Antibiotic resistance spread in food. Nature 389:801–802.
35. Petsaris, O., F. Miszczak, M. Gicquel-Bruneau, A. Perrin-Guyomard, F.
12. del Solar, G., P. Acebo, and M. Espinosa. 1995. Replication control of
Humbert, P. Sanders, and R. Leclercq. 2005. Combined antimicrobial resis-
plasmid pLS1: efficient regulation of plasmid copy number is exerted by the
tance in Enterococcus faecium isolated from chickens. Appl. Environ. Mi-
combined action of two plasmid components, CopG and RNA II. Mol.
36. Roberts, A. P., P. A. Johanesen, D. Lyras, P. Mullany, and J. I. Rood. 2001.
13. del Solar, G., R. Giraldo, M. J. Ruiz-Echevarria, M. Espinosa, and R.
Comparison of Tn5397 from Clostridium difficile, Tn916 from Enterococcus
Diaz-Orejas. 1998. Replication and control of circular bacterial plasmids.
faecalis and the CW459tet(M) element from Clostridium perfringens shows
Microbiol. Mol. Biol. Rev. 62:434–464.
that they have similar conjugation regions but different insertion and excision
14. Doyle, M. P., and M. C. Erickson. 2006. Emerging microbiological food
safety issues related to meat. Meat Sci. 74:98–112.
37. Roberts, M. C. 2005. Update on acquired tetracycline resistance genes.
15. European Commission. 2001. Opinion of the Scientific Committee on Ani-
FEMS Microbiol. Lett. 245:195–203.
mal Nutrition on the criteria for assessing the safety of micro-organisms
38. Ross, J. I., E. A. Eady, J. H. Cove, and W. J. Cunliffe. 1998. 16S rRNA
resistant to antibiotics of human clinical and veterinary importance. http:
mutation associated with tetracycline resistance in a gram-positive bacte-
//www.europa.eu.int/comm/food/fs/sc/scan/out64_en.pdf. Revised 18 April
rium. Antimicrob. Agents Chemother. 42:1702–1705.
39. Sapkota, A. R., K. K. Ojo, M. C. Roberts, and K. J. Schwab. 2006. Antibiotic
16. Florez, A. B., M. S. Ammor, S. Delgado, and B. Mayo. 2006. Molecular
resistance genes in multidrug-resistant Enterococcus spp. and Streptococcus
analysis of a chromosomally encoded erm(B) gene and its flanking insertion
spp. recovered from the indoor air of a large-scale swine-feeding operation.
points in Lactobacillus johnsonii G41. Antimicrob. Agents Chemother. 50:
Lett. Appl. Microbiol. 43:534–540.
40. Speer, B. S., L. Bedzyk, and A. A. Salyers. 1991. Evidence that a novel
17. Gevers, D., G. Huys, and J. Swings. 2003. In vitro conjugal transfer of
tetracycline resistance gene found on two Bacteroides transposons encodes
tetracycline resistance from Lactobacillus isolates to other Gram-positive
an NADP-requiring oxidoreductase. J. Bacteriol. 173:176–183.
bacteria. FEMS Microbiol. Lett. 225:125–130.
41. Tettelin, H., V. Masignani, M. J. Cieslewicz, C. Donati, D. Medini, N. L.
18. Gevers, D., G. Huys, F. Devlieghere, M. Uyttendaele, J. Debevere, and J.
Ward, S. V. Angiuoli, J. Crabtree, A. L. Jones, A. S. Durkin, R. T. Deboy,
Swings. 2000. Isolation and identification of tetracycline resistant lactic acid
T. M. Davidsen, M. Mora, M. Scarselli, Y. Margarit, I. Ros, J. D. Peterson,
bacteria from pre-packed sliced meat products. Syst. Appl. Microbiol. 23:
C. R. Hauser, J. P. Sundaram, W. C. Nelson, R. Madupu, L. M. Brinkac,
R. J. Dodson, M. J. Rosovitz, S. A. Sullivan, S. C. Daugherty, D. H. Haft, J.
19. Gevers, D., L. Masco, L. Baert, G. Huys, J. Debevere, and J. Swings. 2003.
Selengut, M. L. Gwinn, L. Zhou, N. Zafar, H. Khouri, D. Radune, G.
Prevalence and diversity of tetracycline resistant lactic acid bacteria and their
Dimitrov, K. Watkins, K. J. B. O'Connor, S. Smith, T. R. Utterback, O.
TETRACYCLINE RESISTANCE IN L. SAKEI Rits 9
White, C. E. Rubens, G. Grandi, L. C. Madoff, D. L. Kasper, J. L. Telford,
44. Wegener, H. C. 2003. Antibiotics in animal feed and their role in resistance
M. R. Wessels, R. Rappuoli, and C. M. Fraser. 2005. Genome analysis of
development. Curr. Opin. Microbiol. 6:439–445.
multiple pathogenic isolates of Streptococcus agalactiae: implications for the
45. Young, J. P., H. L. Downer, and B. D. Eardly. 1991. Phylogeny of the
microbial ‘pan-genome. ' Proc. Natl. Acad. Sci. USA 102:13950–13955.
phototrophic rhizobium strain BTAi1 by polymerase chain reaction-based
42. Teuber, M., L. Meile, and F. Schwarz. 1999. Acquired antibiotic resistance in
sequencing of a 16S rRNA gene segment. J. Bacteriol. 173:2271–
lactic acid bacteria from food. Antonie van Leeuwenhoek 76:115–137.
43. van Hoek, A. H. A. M., I. M. Scholtens, A. Cloeckaert, and H. J. M. Aarts.
46. Z'Graggen, W. J., H. Fankhauser, F. Lammer, T. Bregenzer, and D. Conen.
2005. Detection of antibiotic resistance genes in different Salmonella sero-
2005. Pancreatic necrosis infection due to Lactobacillus paracasei in an im-
vars by oligonucleotide microarray analysis. J. Microbiol. Methods 62:13–23.
munocompetent patient. Pancreatology 5:108–109.
Source: http://edepot.wur.nl/20972
Wiri Wai Care Wonders! Students from Wiri Central School proudly sign W after completing work on the Puhinui stream. Learning goes beyond the classroom Drury School drain painting Wai Care Fieldtrip to Hunua Brendon with a longfin eel found during fish monitoring Drury School's Wai Care club recently painted up a Papakura Normal School's middle and senior
Osteoporos Int (1997) 7:390–406 ß 1997 European Foundation for Osteoporosis and the National Osteoporosis Foundation Position Paper Guidelines for Diagnosis and Management of Osteoporosis J. A. Kanis, P. Delmas, P. Burckhardt, C. Cooper and D. Torgerson on behalf of the European Foundation for Osteoporosis and Bone Disease Preamble. Significant developments have occurred in





