Microsoft word - 7-4-11-25.docx


International Journal of Applied Research in Natural Products
Vol. 7 (4), pp. 11-25.
Directory of Open Access Journals
2008-2014. IJARNP-HS Publication
Original Research
Extraction and characterization of candidate bioactive
compounds in different tissues from salmon (Salmo salar)
Falkenberg SS*1, Mikalsen S-O2, Joensen H2, Stagsted J3, Nielsen HH1
1Technical University of Denmark, National Food Institute, Søltofts Plads bldg. 221, 2800 Kgs. Lyngby, Denmark
2University of the Faroe Islands, Department of Science and Technology, Nóatún 3 FO-100 Tórshavn, the Faroe Islands
3Department of Food Science, Aarhus University, Blichers Allé 20, 8830 Tjele, Denmark
Summary. There is an interest in bioprospecting organisms from the aquatic environment to find novel bioactive compounds with health
promoting or other functional properties. The aim of this study was to evaluate extracts from untreated and heat-treated salmon tissues for their
radical scavenging activities and for their ability to inhibit activity of the proteases angiotensin I-converting enzyme (ACE) and dipeptidyl
peptidase 4 (DPP-4). In vitro assays were used to detect these activities and the corresponding candidate bioactive compounds were characterized
by LC-MS/MS.
Radical scavenging activity was detected in <10kDa extracts of gills, belly flap muscle and skin with EC50 values of 39, 82 and 100 µg/mL,
respectively. No ACE or DPP-4 inhibiting activity could be detected. LC-MS/MS analysis of dominating compounds in active fractions from size
exclusion chromatography showed that families of related compounds were found in several fractions from different tissues but most pronounced
in gills. One family was defined according to content of a specific amino acid sequence (PW). Three families were defined by the m/z value of
the smallest compound reported in each family (219, 434 and 403). The three latter families did not contain standard unmodified amino acids,
indicating peptides with modified amino acids or other kinds of molecules.
Industrial relevance. Bioprospecting in fish tissue traditionally regarded as waste can lead to detection of novel natural bioactive compounds
including peptides, which could have nutritional, pharmaceutical or other functional value and be used in health and functional foods, thus
increasing the value adding of secondary marine products. A number of naturally occurring antimicrobial peptides have been characterized from
fish skin and gills, such as piscidins, but these and other fish tissues may contain numerous other compounds with bioactive properties. Such
compounds could be extracted by the subsection of the fish industry that processes marine secondary products and further developed to
commercial products. Thus, the identification of novel bioactive compounds could be utilized by the pharmaceutical and biotech industry to
develop new products.
Keywords. Salmon tissue; natural compounds; radical scavenging; marine by-products
INTRODUCTION
Fish are adapted to living conditions utterly different from land-living organisms. For instance, fish must in general possess an
eminent bioactive defense to protect them from the high bacterial load in water, and may thus be expected to harbor bioactive molecules for this purpose (Harnedy & FitzGerald 2012). Bioprospecting in fish tissues that traditionally has been regarded as waste, or has been used for low-value purposes, like animal feed, can lead to detection of novel natural bioactive compounds including peptides with health promoting and functional properties.
A large number of natural antimicrobial peptides (AMPs) are found in invertebrates, and such peptides have also been
identified in a number of fish species (Rajanbabu & Chen 2011). Some AMP families are unique for fish such as the antimicrobial
polypeptides piscidins from gills (Corrales et al., 2009, Corrales et al., 2010). Studies have shown that some of these peptides also exhibit immunomodulatory and antitumor activity (Rajanbabu & Chen 2011). It is therefore possible that some of these peptides are multifunctional and may exhibit other highly relevant bioactivities with health promoting properties eg., oxidative stress that can be suppressed by radical scavenging molecules (Katayama & Mine 2007), hypertension that can be reduced by angiotensin I-converting enzyme (ACE) inhibiting molecules (Fyhrquist & Saijonmaa 2008) and insulin secretion that can be influenced by dipeptidyl peptidase IV (DPP-4) inhibiting molecules (Flatt et al., 2008). The past two decades, the identification of possibly bioactive compounds, present in marine secondary products, has been an emerging area of research (Kim and Mendis 2006). Several investigations have shown that processing leftovers contain significant levels of proteins, cryptides and peptides with bio-functional and techno-functional properties. Peptides derived from marine proteins (cryptides) have proved to be effective against several ailments, such as hypertension (Enari et al. 2008, Gu et al. 2011), osteoporosis (Kanis 2002), cardiovascular diseases (Wergeland et al. 2004), and cancer (Picot et al. 2006). However, like mammals, fish tissue also contains, in variable amounts, some peptides or amino acids with antioxidative activity such as the imidazoles anserine, carnosine and histidine (Abe et al., 1983; Shirai et al., 1983; Suzuki et al., 1990) and glutathione (Bauchart et al., 2007). Additionally, one study has shown that anserine possesses angiotensin converting enzyme (ACE) inhibiting properties (Hou et al., 2003).
_ *Corresponding Author. [email protected] +45 288 02 773 Available online http.//www.ijarnp.org
Falkenberg et al.
Recently, Pampanin et al. (2012) compared peptides with known bioactivities, such as ACE inhibition, antioxidative, or
immunomodulatory activities, with small peptides from herring tissues detected by LC-MS/MS. Based on their sequences, a number of herring peptides with potential bioactivities were suggested. None of the peptides appeared to contain modified amino acids. However, a review by Harnedy & Fitzgerald (2013) shows salmon and salmon by-products, compared to many other marine organisms, have not been subjected to the same extent of bio-prospective multifaceted research activities.
Furthermore, only few studies have isolated and characterised novel naturally occurring low molecular compounds in fish
tissue that can exhibit radical scavenging activity or act as inhibitors toward ACE. To our knowledge studies investigating compounds from fish tissues with DPP-4 inhibiting properties have not previously been carried out. Farmed salmon is nutritional and healthy and has become a popular food worldwide. The market is not satiated yet and the scale of production is still increasing. According to FAO FishStat, the global aquaculture annual production of Salmo salar (Linnaeus, 1758) in 2012 was approx. 2 million tons. The remaining processing leftovers from salmon production amounting to hundreds of thousand tons per annum include trimmings, frames, fins, heads, skin, shells and viscera. These secondary products are normally used for production of fish oil, essential fatty acids, fishmeal, fish silage and animal feed, but can also be utilized for production of high-priced products. The extraction and subsequent exploitation of marine by-products for components with bioactive properties represents an excellent strategy for added-value generation.
The aim of this study is to evaluate radical scavenging activity, ACE and DPP-4 inhibiting activities in extracts from skin, belly
flap muscle and gills from salmon, and characterize the corresponding candidate bioactive compounds to observed activities.
MATERIALS AND METHODS
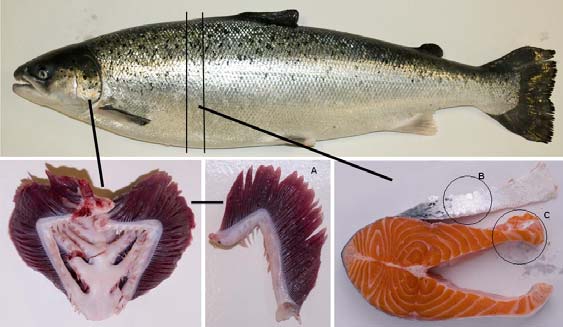
Salmon tissues. Gills, skin and belly flap muscle from fresh Atlantic salmon (Salmo salar) (Fig. 1) obtained from a
commercial fish farm were vacuum packed and stored at - 40°C until use.
Preparation of tissue extracts with and without heat treatment. Thawed gills, skin and belly flap muscle from salmon were
cut into small pieces (approx. 0.5x0.5x0.5 cm for gill and belly flap muscle and 0.5x0.5x0.1cm for skin). For each tissue two samples were prepared where tissue pieces were mixed with 3 volumes of water (w/v) (e.g. 20 g fish tissue + 60 mL water). One sample was boiled in a water bath at 100°C for 10 min. The other sample was not boiled but kept on ice for 10 min. After heating, the tissue/water suspensions were homogenized with an ultraturrax T25 (IKA Labortechnich) at 13500 rpm for 5 min on ice. The suspensions were centrifuged for 45 min at 21000xg at 4°C. The supernatant was filtered through a 150 mm filter paper (Frisenette nr. 201), followed by filtration through 0.45 and 0.20 µm filters (Sartorius) on ice. The obtained filtered extracts were kept on ice until ultra filtration.
Chemicals. 2,2'-azino-bis(3-ethylbenzothiazoline-6-sulphonic acid) (ABTS), Gly–Pro-p-Nitroanilide, captopril, 1-methyl-L-
tryptophan, 5-methyl-DL-tryptophan, diprotin and protease type XIV from Streptomyces griseus ≥ 3,5 units pr mg (pronase) were purchased from Sigma Aldrich (St. Louis, MO, USA). Abz-Gly-Phe(NO2)-Pro was purchased from Bachem AG
(Bubendorf, Switzerland).
Preparation of mucosal extract. Mucosal extract from pig intestine was used as enzyme source for DPP-4 and ACE activity.
Pig intestine was obtained from a Danish landrace/ Yorkshire cross that had been fasted overnight before slaughter and then stored at - 20°C. Jejunum (1 m) was filled with 100 mL of 0.1 M Tris-HCl buffer, pH= 8.0 (22˚C) and inverted briefly 5 times. The crude extract was filtered through a fine-meshed sieve and centrifuged at 4000xg for 30 minutes at 4˚C. The supernatant was saved and 40 mL was dialysed for 24 h at 2˚C in 10 kDa dialysis tubing against 100 x volume of 0.1 M Tris-HCl buffer pH 8. Protein content of the recovered dialysed mucosal extract was 7.2 mg/mL as determined with Pierce BCA protein assay (Thermo Scientific, Rockford, IL, USA) using bovine serum albumin (BSA) as standard. The recovered dialysed mucosal extract was frozen in aliquots at -20˚C until use.
Ultrafiltration of extracts. The filtered extracts were fractionated by ultrafiltration with Vivaspin 10 kDa cut off filters
(Sartorius) at 4000xg at 4°C for 45 min. The filtrates named as <10 kDa extracts were stored at -20°C until use.
Peptide content. Peptide content in the <10 kDa extracts was measured with the Pierce Modified Lowry Assay (Thermo Fisher
Scientific, Rockford, IL, USA). The lower peptide content in the fractions obtained from size exclusion chromatography was measured with the more sensitive Pierce BCA Protein Assay. For both methods BSA was used as standard and peptide content is expressed as mg protein per mL.
ABTS+ radical scavenging activity (modified after Clausen et al., 2009). A volume (50 µl) of <10 kDa extracts or dilutions
thereof were combined in 96 well microplates with 200 µl of ABTS+ solution (0.37 mM in 0.1 M borate buffer, pH=8.0), which had been prepared from 18.7 mM ABTS and 8.8 mM ammonium persulfate in water that had been incubated overnight at room temperature. All dilutions were done in borate buffer. Absorbance was measured at 734 nm in a microplate reader (Biotek, Synergy 2). ABTS+ free radical scavenging activity was defined as decrease in absorbance after 30 min. 50 µl of water was used as control for 100% absorbance.
DPP-4 inhibitory activity (modified after Li-Chan et al., 2012). A volume (50 µl) of <10 kDa extracts or dilutions thereof
were combined with 50 µl of intestinal mucosal extract (diluted 27 times in 100 mM Tris-HCl buffer pH=8.0) in 96 well microplates and prewarmed to 37˚C. All dilutions were done in 0.1 M Tris-HCl buffer, pH 8.0, 25°C. Prewarmed DPP-4 substrate (200 µl of 2.5 mM Gly-Pro-Nitroanilide, 0.1 M Tris-HCl buffer, pH 8.0, 25°C) was added and absorbance measured at 405 nm (and 600 nm to correct for light scatter) every two minutes for 20 minutes in a microplate reader (Synergy 2, Biotek) at 37°C. 50 µl of water was used as control for 100% activity and 2 mM diprotin was used as reference inhibitor. DPP-4 activity was calculated as ∆ mOD 405 nm/min as initial rate for the linear part of the curve and scatter subtracted if present. Inhibition was calculated as activity in % of activity in control.

Bioactive compounds in salmon tissue
ACE inhibitory activity (modified after Sentrandreu & Toldra 2006). A volume of 50µl of <10 kDa extracts in 3-fold
dilution was added to a well in 96 well microplate. Intestinal mucosal extract (50 µl of 10 times diluted extract in 150 mM Tris-HCl buffer pH 8.3) was added to the wells. Before addition of substrate, the microplate was preincubated at 37˚C for 10 minutes in the fluorescence spectrophotometer (Gemini max, Molecular Devices) with automatic mixing. The substrate working solution (light sensitive) (0.45 mM Abz-Gly-Phe(NO2)-Pro in 150 mM Tris-HCl buffer pH 8.3, 25°C with 1.125 M NaCl) was also
incubated at 37˚C for 10 minutes in a water bath. Subsequently, 200 µl of substrate solution was added. 50 µl of water was used as control for 100% activity and 0.12 µM captopril was used as reference inhibitor. Fluorescence was measured every minute as Relative Fluorescence Units (RFU) after automatic shaking for a total of 40 minutes at λ (excitation) = 355 nm and λ (emission) = 405 nm. ACE activity was defined as ∆ RFU per min and inhibition was calculated as activity in % of activity in control.
Figure 1. Picture of the salmon tissues used for extraction of bioactive compounds. (A) Gill, (B) Skin, ( C) Belly flap muscle
Experimental design
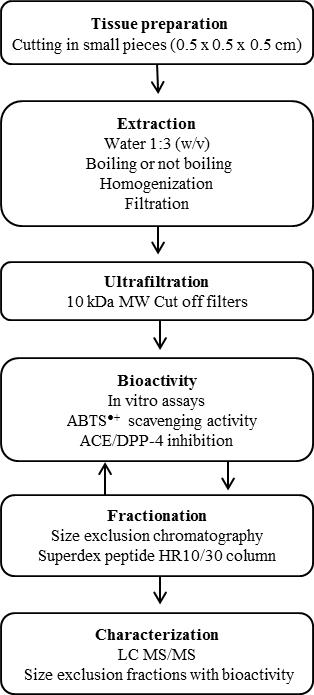
Figure 2. Schematic flowchart showing the experimental design of analyzing bioactive compounds in tissues from salmon
Determination of EC50 values. EC50 values were determined by dose-response experiments, where ABTS+ radical scavenging
and ACE and DPP-4 inhibitory activity were measured with 3-fold dilutions of extracts. Non-linear regression for estimation of EC50 values was performed using sigmoidal dose-response curves with individual Hillslope coefficients (y=min+ (max-
min)/(1+10 (EC50-x)Hillslope) built into Sigmaplot v. 11 graphing and statistics software (Systat software Inc)
Falkenberg et al.
Size exclusion chromatography (SEC). Compounds in <10 kDa extracts were separated by FPLC (Fast Performance Liquid
Chromatography) using an Äkta Purifier system with FRAC 950 collector. Undiluted filtrate (200 µl) was injected into a SuperdexTM peptide 10/300 GL column (GE Healthcare) using 100 mM ammonium acetate buffer pH 8 with a flow rate at 0.5 mL/min. Ammonium acetate is a volatile buffer that minimally affects subsequent steps of mass spectrometry. Compounds were detected at 215 nm and 280 nm. Fractions of 1 mL each were collected (35 fractions in total) and stored at -20°C until use. Cytochrome c (12.3 kDa), aprotinin (6.5 kDa), Lys5 (659 Da), Gly3 (189 Da), and Gly (75 Da) (Sigma-Aldrich, St. Louis,
Missouri, USA) were used as molecular weight markers.
Pronase treatment of extracts. Pronase (1 mg/mL) dissolved in 100 mM Tris-HCl pH 8 was mixed with <10 kDa extracts in
the ratio 120 µl to 480 µl (pronase 5 times dilution, Enzyme:Substrate ratio 1:4 (v:v)) in a microcentrifuge tube. Control samples without pronase but with water were made the same way. A sample containing only pronase (1 mg/mL) was made to indicate elution of pronase and corresponding autolytic peptides, if any. The microcentrifuge tubes were placed in a 37˚C water bath overnight (16 hours) and immediately analysed by SEC as described above.
LCMS and LCMS/MS. Fractions from the size exclusion chromatography that showed antioxidative capacity were dried in a
SpeedVac (Thermo Scientific Savant SPD1010) and reconstituted in 50 µl buffer A (5% acetonitrile [ACN] + 0.1% formic acid [FA]), vortexed, centrifuged at room temperature at 14,000xg for 10 min and transferred to LC vials. Buffer B was 90% ACN + 0.1% FA. A nanoHPLC (EASY nLC; Proxeon, Odense, Denmark) was used. The samples were trapped on an EASY reverse phase pre-column (2 cm length, ID 100 µm, 5 µm C18 beads) and separated on an EASY reverse phase analytical column (10 cm length, ID 75 µm, 3 µm C18 beads). Flow rate was 300 nL/min. The time from mixer to elution was 3 min, corresponding to a dead volume of around 900 nL. The components were eluted by a gradient (0 min 100%A, 0.05 min 99%A, 2 min 90%A, 21 min 45%A, 23 to 26 min 0%A, 28 min 95%A, 30 min 100%A) into a micrOTOF-QII mass spectrometer (Bruker Daltonik, Bremen, Germany) through the standard nanosprayer into the electrospray ion source. Nitrogen was used as nebulizer (pressure at 1 bar) and drying gas (5 L/min at 150 °C) in the ion source. Argon 5.0 was used as collision gas, and the collision energy was automatically adjusted to the selected m/z values using default options. Infusion of electrospray calibrant solution (Fluka) by a syringe pump was used as external calibration for the samples. Additionally, a standard mixture of tryptic peptides made from BSA (Agilent) was run for every fifth LC-MS/MS sample, and was used for more detailed external calibration. The MS instrument was controlled by instrument-specific software (Compass 1.3 micrOTOF control v2.3), where the acquisition parameters were set (positive mode, low mass limit at m/z 50, MS/MS auto in the m/z range 70 to 800, 4 precursor ions with absolute intensity > 2000 counts, exclude after 4 spectra, release after 1 min). The data were initially analyzed in the Compass 1.3 DataAnalysis v4.0 software to generate the compounds and externally calibrate the spectra. The software extracted compounds using intensity threshold at 1000, retention time window 1 min, S/N threshold 3, and relative area and intensity thresholds at 3%.
RESULTS AND DISCUSSION
In the last decade a large number of small antimicrobial peptides have been identified in fish tissue like gills, skin and organs
(Hoang & Kim, 2013). Studies have shown that some of these peptides also can have other bioactive properties like anticancer activity (Chang et al., 2011) and immunomodulatory activity (Hsieh et al., 2010). This suggests that salmon tissue could have unidentified endogenous peptides or other compounds that possess other bioactive properties apart from antimicrobial activity. This is supported by the study by Pampanin et al. (2013), who showed that extracts from herring skin and internal organs contained many small peptides with potential bioactive properties. The present study analysed three different tissues from salmon for low molecular weight compounds (<10 kDa) with radical scavenging activity and ACE and DPP-4 inhibitory properties based on the hypothesis that salmon tissue contains novel bioactive compounds. Extraction of compounds was done in water with heat treatment to prevent endogenous proteolysis and also to prevent bacterial growth. However as heat treatment may also affect relevant compounds such as peptides, an extraction was also carried out without heat treatment. Analysis with Lowry assay showed that boiling did not affect the peptide content in <10 kDa extract from gill compared to extract prepared without boiling (Table 1). An increase in the amount of recovered peptides was observed in both belly flap muscle and skin extract after boiling (Table 1), which could be due to a more efficient extraction from these tissues.
Table 1. Peptide contents and radical scavenging activity in <10 kDa extracts of different tissues from salmon. Peptide content was measured in
duplicates by the Lowry method using bovine serum albumin (BSA) as standard, and is given as mg protein/mL. Radical scavenging activities are
reported as the peptide concentrations that were able to remove 50% of the ABTS+ radicals (EC50, half maximal effective concentration). EC50
values were estimated by non-linear regression (sigmoidal dose-response curve) with 95% confidence intervals.
Treatment
EC50 (mg/mL) EC50 (mg/mL) 95%
confidence interval
2.9 x 10-3 - 4.2 x 10-3
Belly flap muscle
9.8 x 10-3 - 1.1 x 10-3
No Boiling
4.6 x 10-3 - 5.7 x 10-3
Belly flap muscle
7.7 x 10-3 - 8.8 x 10-3
9.9 x 10-3 - 1.6 x 10-2
Bioactive compounds in salmon tissue
Bioactivities. All <10 kDa extracts showed clear ABTS+ radical scavenging activities with 95% decrease in absorbance at
734 nm (Fig. 3). Dose-response data from analyses of three-fold dilutions of the extracts are shown in Fig. 3. There were relatively small differences between the tissues, with gill extracts apparently giving the lowest EC50 value estimated by non-linear
regression analysis (Table 1), which may suggest that this extract contained compounds with more potent radical scavenging
activity than extracts from either belly flap muscle or skin.
Figure 3. Concentration dependencies of ABTS+ radical scavenging activities in extracts (<10 kDa) from salmon tissues. Extracts were from (A)
boiled tissues (filled symbols) or from (B) control (not boiled) tissues (open symbols). The tissues were gill (,○); belly flap muscle (■,□ ) and
skin (▲,∆). The curves were fitted by nonlinear regression – global curve fitting. Absorbance was normalized to 1 based on absorbance of water
as control.
None of the <10 kDa extracts inhibited DPP-4 and ACE activity when compared to water as control (results not shown). This is
notable as Pampanin et al. (2012) found that extracts of herring skin with PBS buffer contained several small peptides with potential bioactivity on the cardiovascular system such as reducing hypertension.
Size exclusion chromatography. Size exclusion chromatography of <10 kDa extracts showed very different elution profiles of
compounds from belly flap muscle, gills, and skin, respectively, as detected by absorbance at 215 and 280 nm (Fig. 4), indicating different compositions of low molecular weight compounds in these tissues. A common feature for boiled extract from all three tissues was that compounds did not elute before 18 mL, which corresponds to the MW of Gly3. This indicated that even though
extracts were fractionated by 10 kDa cut-off ultrafiltration, the largest compounds eluting were in the range of 400-500 Da assuming that there was no extraneous interaction between the compounds and column matrix. Both at 280 nm and 215 nm between 3 and 5 peaks eluted after Gly. This indicates that all extracts contained compounds that adsorbed to the column and therefore could have an MW larger than Gly. Only control extracts from gills contained compounds that eluted with void volume, which suggested a MW larger than 7 kDa. These compounds could have precipitated during boiling as they were not detected in the filtrate. Boiling also influenced the profile of belly flap muscle as shown by an increase in compounds eluting between 20 and 22 ml. In contrast no effect of boiling was observed for skin.
Falkenberg et al.
Not Boiled
Belly flap muscle
Elution volume (ml)
Elution volume (ml)
Figure 4. Size exclusion chromatography (SuperdexTM peptide 10/300 GL column) of <10 kDa extracts (boiled and not boiled) from three
different salmon tissues: gills (top row), belly flap muscle (middle row) and skin (bottom row). Designation of peptide standards markers is
shown on figure "Gill" (top row, left column). Eluting components were detected by absorbance at 215 and 280 nm.
Test of peptide content in peaks from size exclusion chromatography. A test of peptide content was made by addition of
pronase to the boiled < 10 kDa extracts and analysed by SEC. Peaks that were affected by the treatment with pronase indicated presence of peptides consisting of standard amino acids.
Bioactive compounds in salmon tissue
Figure 5. Effect of pronase (Streptomyces griseus protease XIV) on Size exclusion chromatography elution profile of < 10 kDa extracts from
boiled salmon tissue. (A) Gills, (B) Belly flap muscle, (C) Skin. ― Without pronase; ······ with pronase. Pronase (1 mg/mL in 100 mM Tris-HCl
pH 8) was mixed with extract in ratio 1:4 (v:v) and incubated 16 hours before analyses by SEC.
Fig. 5 shows the elution profiles of < 10 kDa extracts from gills, belly flap muscle and skin with and without pronase. Pronase
eluted as a single peak at 9 mL corresponding to the void volume. In general, the pronase treatment had surprisingly little effect on the extracts. The resistance to proteolysis indicated the presence of compounds other than unmodified peptides. The main effect of pronase on the gill extract was on a peak eluting at 22 mL, which was reduced considerably, while a peak eluting at 24 mL correspondingly increased (Fig. 5A). The pronase treatment on belly flap muscle extract (Fig. 3B) mainly resulted in a decrease of a major peak eluting at 19 mL. An increase in a peak eluting at 19.5 mL could be a result of cleavage products from peptides eluting at 19 mL. Three peaks eluting at 20.5, 22 and 26 mL also decreased after pronase treatment. Interestingly, the decrease of these peaks did not result in increase in new peaks eluting later, possibly because pronase destroyed the peptide bonds. The effect of pronase treatment on skin extract seemed somewhat smaller than on gill and belly flap muscle extract (Fig. 5C). As for belly flap muscle extract, a decrease in peaks eluting at 19 and 26 mL was seen.
Falkenberg et al.
ABTS+ radical scavenging activity and peptide content in fractions from boiled salmon. Peptide content and radical
scavenging activities were measured in fractions from the size exclusion chromatographed samples of boiled < 10 kDa extracts from salmon gills, belly flap muscle and skin (Fig. 6). There was no ABTS+ scavenging activity in fractions eluting from 0-16mL. In the fraction eluting at 18 mL, a weak ABTS+ radical scavenging activity was observed for gills and skin, which could be due to peptides in the low molecular weight range (between Gly3 and Lys5). Pampanin et al. (2013) identified small
peptides in extracts from herring skin that could have antioxidative properties, suggesting that fish skin in general contains small
peptides with antioxidative properties. However, there is not a full correlation between peptide content and radical scavenging
activity indicating the presence of inactive peptides or that other types of compounds were contributing to the activity. The
strongest ABTS+ radical scavenging activity was detected in fractions eluting between 21 and 22 mL for all three tissues with
strongest activity in the gill extract, where also the highest peptide content was detected. This is in agreement with the lower EC50
value for gill extract compared to the two other tissue extracts. These fractions eluted after glycine and suggest the presence of compounds with some affinity to the column material. Radical scavenging activity was also detected in fractions eluting at 28 and 33 mL for gill and skin, respectively, indicating even stronger interaction with the column. Fractions from untreated salmon extracts were also analyzed with the ABTS+ radical scavenging assay, and the activity profiles matched those from boiled salmon extract (results not shown).
Figure 6. Peptide content (A) and ABTS+ radical scavenging activity (B) in fractions from size exclusion chromatography of <10 kDa extract
from boiled salmon tissue. () Gills,( ○) Belly flap muscle, (+) Skin. Peptide standards markers are shown in A. Peptide content was analysed
with bicinchoninic acid assay (Pierce BCA Protein Assay) with bovine serum albumin as standard.
LC-MS and MS/MS. The fractions from the boiled samples that showed the highest ABTS+radical scavenging activity were
selected for analysis by mass spectrometry. Focus was on the major compounds in the fractions (Fig. 5a, b and c) that gave MS/MS spectra of reasonable quality. C18 reverse phase columns were used in the LC-MS analyses. Very hydrophilic compounds are not retained on such columns, and will therefore not be detected.
Bioactive compounds in salmon tissue
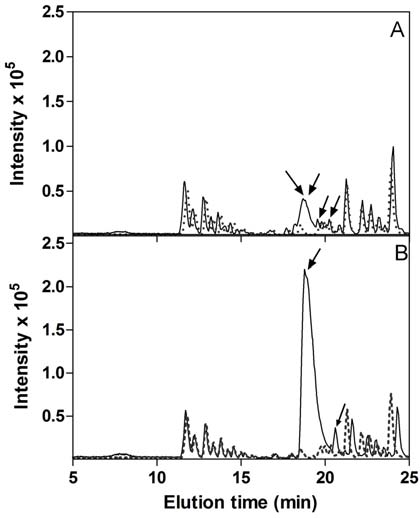
Figure 7a. Base peak ion chromatograms from nanoLC reverse phase separation of fractions from salmon gill. The fractions were from size
exclusion chromatography (SEC) performed on <10 kDa extract from salmon gill, and collected in 1 ml portions at (A) 18 mL (B) 21 mL, (C) 22
mL, (D) 28 mL, (E) 29 mL (cf. Fig 4), corresponding to the fractions showing the highest ABTS+ radical scavenging activity. ― Sample; ---
Blank; Arrows indicate peaks containing components that have been characterized by MS/MS and are mentioned in text and tables.
Falkenberg et al.
Figure 7b. Base peak ion chromatograms from nanoLC reverse phase separation of fractions from belly flap muscle. The fractions were from
size exclusion chromatography (SEC) performed on <10 kDa extract from salmon belly flap muscle, and collected in 1 ml portions at (A): 21 mL,
(B) 22 mL. (cf. Fig 4), corresponding to the fractions showing the highest ABTS+ radical scavenging activity. ― Sample; --- Blank; Arrows
indicate peaks containing components that have been characterized by MS/MS and are mentioned in text and tables.
Figure 7c. Base peak ion chromatograms from nanoLC reverse phase separation of fractions from salmon skin. The fractions were from size
exclusion chromatography (SEC) performed on <10 kDa extract from salmon skin, and collected in 1 ml portions at 22 mL (cf. Fig 4),
corresponding to the fraction showing the highest ABTS+ radical scavenging activity. ― Sample; --- Blank; Arrow indicate peak containing
components that have been characterized by MS/MS and are mentioned in text and tables.
The mass spectrometric analyses showed three conspicuous features. First, some compounds were found in several fractions
and from different tissues. Second, several families of chemically related compounds were evident. In fact, most compounds belonged to one or another multimember family, here called the "PW family", "434 family" "403 family" and "219 family", as recognized by similarities of the MS/MS spectra within each family (Tables 2, 3, 4 and 5). The families are named according to a specific amino acid sequence (PW) or by the lowest m/z value reported in each family. Third, there were only few compounds with standard amino acids as major constituents. No proteolytic treatment was performed on the fractions analyzed by MS and endogenous proteases were inactivated by the heat treatment. Thus, the detected compounds were likely small compounds or peptides freely present in the tissues, although the boiling procedure may have resulted in heat-induced chemical modifications.
The only compounds that consisted of identifiable amino acids were found in fraction 21 and to a lesser degree in fraction 22
from gills. M/z 399 (Fig. 8) could be explained as the peptide PPW (theoretical m/z 399.203, observed m/z 399.202), member of the PW family (Table 2). The compound m/z 486 was apparently related to m/z 399 (both having MS/MS fragments at m/z 188, 205 and 302), although the former had a somewhat poor MS/MS spectrum. The mass difference between 399 and 486 could suggest an additional Ser, thus giving the sequence SPPW or PSPW (theoretical m/z 486.235, observed m/z 486.235). Comparing with zebra fish sequences (the salmon genome is not yet available), the PPW sequence is found in numerous proteins, but we are not aware of any previous reports indicating that it may be present as a free peptide. Also the sequences SPPW or PSPW can be
Bioactive compounds in salmon tissue
found in several tens of zebra fish proteins. The PPW and SPPW/PSPW could be responsible for part of the radical scavenging
activity observed in fraction 21 of gill and could represent new antioxidative peptides naturally present in fish tissue.
Table 2. Two compounds in the PW family and two unrelated compounds detected by reverse phase LC-MS/MS. These compounds were only
found in <10 kDa extract from salmon gills.
Obs. m/z, charge
146.06, 159.09, 188.07, 205.10, 302.15
Gills fraction 21
PPW (theoretical m/z 399.20). PW family.
157.10, 188.08, 205.10, 302.15
Gills fraction 21
(theoretical m/z 486.24). PW family
132.08, 141.10, 144.08, 159.09, 170.06, Gills fractions 21 and 22
(theoretical m/z 417.25)
125.02, 141.08, 143.03, 152.07, 183.04, Gills fraction 28
Table 3. Compounds in the m/z 434 family detected by reverse phase LC-MS/MS. These compounds were only found in <10 kDa extract from
salmon gills.
Obs. m/z, charge
116.02, 129.07, 133.10, 141.07, 142.03, Gills fraction 18
144.01, 155.09, 162.02, 167.09, 170.03, 185.10, 199.07, 233.06
116.02, 129.07, 133.10, 142.03, 144.01, Gills fraction 18
157.10, 162.02, 170.03, 187.11, 199.07, 233.07
116.02,127.06, 129.07, 142.03, 144.01, Gills fraction 18
155.09, 162.03, 170.03, 173.10, 199.11, 216.14, 233.06
Table 4. Compounds in the m/z 403 family detected by reverse phase LC-MS/MS. These compounds were only found in <10 kDa extract from
salmon gills.
Obs. m/z, charge
112.05, 119.04, 136.06, 162.07, 176.00, Gills fraction 21
Likely the same as the co-
193.05, 250.10, 288.05, 306.05, 348.07,
eluting 604; here as (M+3H+)/3
112.05, 119.04, 136.06, 152.06, 161.06, Gills fraction 22
192.08, 216.09, 290.06, 332.08
119.03, 135.03, 152.06, 161.06, 216.09, Gills fraction 21
234.10, 314.07, 33208, 394.10
119.03, 135.04, 136.06, 152.06, 161.06, Gills fraction 21
216.09, 232.09, 312.05, 332.08, 348.09
112.05, 119.03, 136.06, 143.04, 152.06, Gills fraction 22
163.05, 176.80, 191.04, 250.09, 312.05, 330.05
112.05, 119.04, 136.06, 152.06, 161.06, Gills fraction 18
Probably different from 455 in
192.08, 216.09, 290.06, 332.07
119.04, 135.04, 136.06, 152.06, 161.06, Gills fraction 22
Probably different from 455 in
216.09, 232.08, 332.07, 348.08
112.05, 119.04, 135.03, 136.06, 152.06, Gills fraction 22
177.00, 193.05, 232.08, 250.10, 312.04, 330.07, 348.07
112.05, 119.03, 136.06, 139.95, 152.06, Gills fraction 18
161.06, 192.08, 216.09, 234.09, 290.06, 314.07, 332.07
112.05, 119.03, 126.07, 135.02, 136.06, Gills fraction 18
152.06, 161.07, 176.99, 192.08, 216.09, 232.08, 314.06, 332.09
112.06, 136.06, 193.05, 208.07, 226.09, Gills fraction 21
Likely the same as the co-
232.08, 250.09, 268.10, 288.05, 291.06,
eluting 403; here as (M+2H+)/2
306.05, 330.06, 348.07, 420.08, 462.11
Falkenberg et al.
Table 5. Compounds in the m/z 219 family detected by reverse phase LC-MS/MS. These compounds were found in all <10 kDa extracts from
salmon tissues (gills, belly flap muscle and skin).
Obs. m/z,
MS(2) fragments
128.06, 133.02, 141.01, 154.08, 159.02, Gills fraction 21, 22, 28, 29
171.03, 173.05, 183,04, 201.05
Belly flap fraction 21, 22 Skin fraction 22
141.01, 159.02, 201.05, 219.06
Gills fraction 21, 22
Belly flap fraction 22
129.07, 133.02, 154.08, 171.04, 173.05, Gills fraction 21
183.04, 201.05, 219.06
141.01, 159.02, 201.05, 219.06
Gills fraction 22
159.02, 173.05, 201.05, 219.06, 259.09
Gills fraction 22
Belly flap fraction 21, 22
141.02, 159.03, 171.03,173.06, 201.05, Gills fraction 28, 29
165.01, 201.04, 219.06, 243.06
Belly flap fraction 21
Figure 8. MS/MS spectrum of compound m/z 399.202 from fraction 21 mL from SEC of gill extract, and its identification as PPW. The MS/MS
fragments with the suggested chemical structures are also generated from free Trp (corresponding to the y1 fragment; data not shown). The m/z
values of these fragments from free Trp are also found in the databases Metlin (metlin.scripps.edu) and MassBank (www.massbank.jp), and together with the Trp-specific y1 fragment at m/z 205, they are probably indicative of a C-terminal Trp. Wi, immonium ion of Trp; P, proline. The small diamond () to the right indicates the m/z value of the intact ion.
A compound of m/z 417 was found in fractions 21 and 22 from gills (Table 2). MS/MS fragments show a strong peak at
159.09, potentially fitting with the immonium ion of Trp. The remaining mass of the original ion could fit with presence of Val and Leu/Ile. Leu/Ile in C-terminal position would give an y1-ion of 132, which is present. Thus, a tripeptide consisting of Trp,
Val, and Leu/Ile, is a possibility (WVL/I or VWL/I), but there are still a number of unexplained fragments left. Thus, this compound is presently considered as unidentified.
All other compounds contained mainly units that were not standard amino acids; i.e., modified amino acids, or some other
kinds of molecules.
The 434 family was found in fraction 18 from gills (Table 3), with compounds of m/z 434, 436 and 448 (distinct from 448 in
the 219 family). The three compounds eluted at different positions in the gradient, and contained a number of fragment ions in common. Additionally, there was one series of fragments that had the same mass difference as the intact ions (m/z 185, 187, 199, respectively).
Bioactive compounds in salmon tissue
The 403 family was found in fractions 18, 21 and 22 from gills (Table 4), and these compounds tended to elute early in the
gradient (8 to 11 min). Most analyzed members in this family were doubly charged (in contrast to all compounds mentioned above). The members were characterized by a strong MS/MS fragment at m/z 136.06 (a value close to the immonium ion of Tyr), and a number of additional common fragments. However, no other data directly supported the presence of Tyr.
Figure 9. MS/MS spectra of representatives in the m/z 219 family. (A) m/z 219. (B) m/z 437. (C) m/z 349. Note the common fragments: m/z
219, 201, 159 in all three spectra, m/z 141 in A and B, and m/z 173 in A and C. The small diamond () to the right indicates the m/z value of the
intact ion.
The 219 family gave the major chromatographic peak(s) in all shown fractions, except in Fig 5a (fraction 18 from gills). The
family consisted of compounds with m/z 219, 261, 293, 349, 437, and 655. These compounds had at least 4 MS/MS fragments in common with m/z 219 (Fig. 9 and Table 5). M/z 219, 437 and 655 were co-eluting, and the latter was only found in fractions where 219 was very intense. They probably corresponded to the same compound (as M + H+, 2M + H+, and 3M + H+, respectively). Also m/z 448 (fraction 21 from belly flap muscle; note that there was also another compound with m/z 448 as indicated below) probably belonged to this family, although it only showed two fragments in common with the remaining family (fragments 219 and 201).
Being the common basis of the most prominent family of compounds found in these experiments, m/z 219 was selected for
further analyses. The isotopic envelope of the intact compound 219 showed intensities of the second and third peaks of 12.84% (mean; range 12.27-13.44%) and 1.17% (mean; range 1.08-1.29%). The chemical composition that most closely approximated the mean isotopic distribution was C10H10N4O2 (as neutral compound; theoretical m/z for MH+ is 219.0877; theoretical peak
intensities of 12.82 and 1.16% as calculated by the NIST08 database software).
There are some biologically relevant compounds that have theoretical m/z values close to the detected in the range 219.05-
219.07. Potassium adducted glucose (theoretical m/z 219.027) can easily be excluded based on both its hydrophilicity and its isotopic envelope distribution. Another candidate could be methylated Trp (theoretical m/z 219.113; chemical composition for neutral compound is C12H14N2O2). If so, this could suggest 261 as the acetylated methyl-Trp. The spectra of 1-methyl-L-Trp and
5-methyl-DL-Trp were therefore investigated. The two methyltrypthophans showed very similar spectra (not shown). Only the MS/MS fragment at 173 was common for the methyltryptophans and compound 219. Furthermore, some of the major fragments showed a 1 Da difference between compound 219 (see also Fig. 9A) and the methyltryptophans (201 vs 202, 183 vs 184, 159 vs
Falkenberg et al.
160, 133 vs 132). Theoretical peak intensities for methylated Trp are 14.35 and 1.35% for second and third peak, and observed intensities were 13.9 and 1.5%, respectively. Thus, the 219 compound is not methyltryptophan, although some kind of chemical relationship cannot be excluded.
The presence of the 219 compound or its family members in most dominating fractions with radical scavenging activity
indicate that these compounds contribute in general to the observed radical scavenging activity in the different tissues. Thus, the 219 compound could be an interesting new natural radical scavenging agent. The 219 compound is further discussed below in the section "Database searches".
Many of the characterized compounds exhibited low fractional masses (the numbers after the decimal point). If the analyzed
compounds consist of the common elements of organic biomolecules, they must contain several rings and unsaturated bonds, in addition to oxygens or phosphorus (sulfur can be excluded based on the isotopic peak distributions). Many bioactive peptides contain amino acids that possess ring and unsaturated bonds. Trp can exhibit radical scavenging activity as the indolic group can serve as hydrogen donor (Pihlanto, 2006). Saito et al. (2003) studied radical scavenging activity of different combinations of tripeptides and found that tripeptides with a Trp at the C-terminus exhibited the highest radical scavenging activity. The presence of Trp could also explain that these peptides elute after Gly when fractionated on the size exclusion column. The aromatic amino acids Phe, Tyr and Trp have been analysed on the SEC column using the previously described method and all three eluted after Gly (results not shown).
Anserine and carnosine, two modified dipeptides with a methylhistidine moiety possessing antioxidant and ACE inhibiting
activities (Hou et al. 2003), have previously been detected in fish tissue (Shirai et al 1983, Bauchart et al 2007). In the studied fractions, we did not detect anserine or carnosine, or any other compound with a methylhistidine moiety.
Database searches. Searches in chemical databases might in some cases be of considerable help in identifying unknown
compounds (Little et al., 2012; Wolf et al., 2010). External calibration with known samples run for every fifth fish sample, gave a range of m/z between 219.0580 and 219.0627 (average 219.0600) for the 219 molecule.
The NIST08 database was searched with the data for m/z 219, but no reasonable hits were found. The compound was then
analyzed by MetFrag (Wolf et al., 2010), using the three databases KEGG, PubChem and ChemSpider in a range from m/z 219.050±25 ppm to 219.100±25 ppm in steps of 0.01, and using the 9 peaks indicated in Table 5. MetFrag suggested several compounds with theoretical fragments covering 5 to 7 of the observed fragments, but when screened against the isotopic distribution, most of the hits could be excluded. Compounds with neutral composition C12H11O2P (e.g., PubChem ID 17841952),
C11H10N2O3 (e.g., PubChem IDs 4268466, 13290790), and C10H10N4O2 (e.g., PubChem ID 28412031) showed 5 to 6 hits with the
observed fragments, and reasonable fits with the isotopic contribution. However, none of them appeared as obvious candidates for being immediately accepted as the presently observed compound, as some of the major peaks were lacking for all hits.
The theoretical isotopic peak intensities may depend somewhat on the software used in the calculations (we have used the
calculator in the NIST08 database software). In principle, the isotopic distribution may slightly change according to food and trophic level of the organism (McCutchan et al., 2003; Newsome et al., 2007; Henry et al., 2012), although it would not be expected that such changes should be detected by the present type of MS. Still, we have compared observed and theoretical peak intensities for some salmon peptides identified in other analyses (Falkenberg et al., in prep). Most of the observed intensities were 92 to 100% of the theoretical intensities, with a few instances (for the third peak in the isotopic envelope) up to 120% of theoretical intensity. Thus, it is possible that the chemical composition with the best calculated fit, C10H10N4O2 for the neutral
compound, could be a close approximation to the real compound with m/z 219.
We thank the Faroese Research Council and The Danish Agency for Science, Technology and Innovation (grant no. 645-08-
0113) for financial support to this study. The mass spectrometry facility at the University of the Faroe Islands is supported by Statoil Faroes, Faroe Islands. We thank P/F Bakkafrost, FO-625 Glyvrar, Faroe Islands for supplying salmon material.
REFERENCES
Abe, H. 1983. Distribution of free L-histidine related dipeptides in the muscle of fresh-water fishes. Comparative Biochemistry and Physiology
Bauchart C, Chambon C, Mirand PP, Savary-Auzeloux I, Remond D, Morzel M. 2007 Peptides in rainbow trout (Oncorhynchus mykiss)
muscle subjected to ice storage and cooking. Food Chemistry 100: 1566-72
Chang WT, Pan CY, Rajanbabu V, Cheng CW, Chen JY. 2011. Tilapia (Oreochromis mossambicus) antimicrobial peptide, hepcidin 1–5, shows
antitumor activity in cancer cells. Peptides 32: 342-352
Clausen MR, Skibsted LH, Stagsted J. 2009. Characterization of major radical scavenger species in bovine milk through size exclusion
chromatography and functional assays. Journal of Agricultural and Food Chemistry 57: 2912-2919
Corrales J, Gordon WL, Nogaa EJ. 2009. Development of an ELISA for quantification of the antimicrobial peptide piscidin 4 and its application
to assess stress in fish. Fish and Shellfish immunology 27: 154-163
Corrales J, Mulero I, Mulero V, Nogaa, EJ. 2010. Detection of antimicrobial peptides related to piscidin 4 in important aquacultured fish.
Developmental and Comparative Immunology 34: 331-343
Enari H, Takahashi Y, Kawarasaki M, Tada M, Tatsuta K. 2008. Identification of angiotensin I-converting enzyme inhibitory peptides derived
from salmon muscle and their antihypertensive effect. Fisheries Science 74: 911-920.
Flatt PR, Bailey CJ, Green BD. 2008. Dipeptidyl peptidase IV (DPP IV) and related molecules in type 2 diabetes. Frontiers in Bioscience 13:
Bioactive compounds in salmon tissue
Fyhrquist F, Saijonmaa O. 2008. Renin-angiotensin system revisited, Journal of Internal Medicine 264: 224–236 Gu RZ, Li CY, Liu WY, Yi WX, Cai MY. 2011. Angitension I-converting enzyme inhibiotory activity of low-molecular-weight peptides from
Atlantic salmon (Salmo salar L.) skin. Food Research International 44: 1536-1540.
Harnedy PA, FitzGerald RJ. 2012. Bioactive peptides from marine processing waste and shellfish: A review. Journal of Functional Foods 4: 6-24 Harnedy PA, FitzGerald RJ. 2013. Bioactive Proteins and Peptides from Macroalgae, Fish, Shellfish and Marine Processing Waste. In Kim S-K,
ed. Marine proteins and peptides: biological activities and applications. John Wiley & Sons Ltd., The Atrium, Southern Gate, Chichester, West Sussex, PO19 8SQ, UK 5-37
Henry AG, Ungar PS, Passey BH, Sponheimer M, Rossouw L, Bamford M et al. 2012. The diet of Australopithecus sediba. Nature 487: 90-93 Hou W-C, Chen H-J, Lin Y-H. 2003. Antioxidant peptides with angiotensin converting enzyme inhibitory activities and applications for
angiotensin converting enzyme purification. Journal of Agricultural and Food Chemistry 51: 1706-1709
Hoang VLT, Kim SK 2013. Antimicrobial peptides from marine sources. Current Protein & Peptide Sciences 14: 205-211 Kanis JA. 2002. Calcitonin in osteoporosis. Bone 30: 65-66. Katayama S, Mine Y. 2007. Antioxidative activity of amino acids on tissue oxidative stress in human intestinal epithelial cell model. Journal of
Agricultural and Food Chemistry 55: 8458–8464
Kim S, Mendis E. 2006. Bioactive compounds from marine processing byproducts – A review. Food Research International 39: 383-393. Li-Chan ECY, Hunag SL, Jao CL, Ho KP, Hsu KC. 2012. Peptides Derived from Atlantic Salmon Skin Gelatin as Dipeptidyl-peptidase IV
Inhibitors. Journal of Agricultural and Food Chemistry 60: 973-978
Little JL, Williams AJ, Pshenichnov A, Tkachenko V. 2012. Identification of "Known Unknowns" Utilizing Accurate Mass Data and
ChemSpider. Journal of the American Society for Mass Spectrometry 23: 179-185
McCutchan JH, Lewis WM, Kendall C, McGrath CC. 2003. Variation in trophic shift for stable isotope ratios of carbon, nitrogen, and sulfur.
Oikos 102: 378–390
Newsome SD, Martinez del Rio, C, Bearhop S, Phillips DL. 2007. A niche for isotopic ecology. Frontiers in Ecology and the Environment 5:
Pampanin DM, Larssen E, Provan F, Sivertsvik M, Ruoff P, Sydnes MO. 2011. Detection of small bioactive peptides from Atlantic herring
(Clupea harengus L.). Peptides 32: 415-420
Picot L, Bordenave S, Didelot S, Fruitier-Arnaudin I, Sannier F, Thorkelsson G, Berge JP, Guerard F, Chabeaud A, Picot JM. 2006.
Antiproliferative activity of fish protein hydrolysates on human breast cancer cell lines. Process Biochemistry 41: 1217-1222.
Pihlanto A. 2006. Antioxidative peptides derived from milk proteins. International Dairy Journal 16: 1306–1314 Rajanbabu V, Chen J-Y. 2011. Applications of antimicrobial peptides from fish and perspectives for the future. Peptides 32: 415-420 Saito K, Jin D-H, Ogawa T, Muramoto K, Hatakeyama E, Yasuhara T, Nokihara K. 2003. Antioxidative Properties of Tripeptide Libraries
Prepared by the Combinatorial Chemistry. Journal of Agricultural and Food Chemistry 51: 3668−3674
Sentandreu MA, Toldra F. 2006. A rapid, simple and sensitive fluorescence method for the assay of angiotensin I-converting enzyme. Food
Chemistry 97: 546-554
Shirai T, Fuke S, Yamaguchi K, Konosu S. 1983. Studies on extractive components of Salmonids - II. Comparison of amino acids and related
compounds in the muscle extracts of four species of salmon. Comparative Biochemistry and Physiology 74B: 685-689
Suzuki T, Hirano T, Shirai T. 1990. Distribution of extractive nitrogenous constituents in white and dark muscles of fresh-water fish.
Comparative Biochemistry and Physiology 96B: 685-689
Wergeland H, Liaset B, Gudbrandsen OA, Lied E, Espe,M, Muna Z, et al. 2004. Fish protein hydrolysates reduces plasma total cholesterol,
increases the proportion of HDL cholesterol, and lowers acyl-CoA:cholesterol acyltransferase in liver of Zucker rats. Journal of Nutrition 134: 1320-1327
Wolf S, Schmidt S, Müller-Hannemann M, Neumann, S. 2010. In silico fragmentation for computer assisted identification of metabolite mass
spectra. BMC Bioinformatics 11:148
Source: http://setur.fo/fileadmin/user_upload/NVD/Svein/Artikler/2014_SusanSkanderupFalkenberg_et_al_2014_IJARNP.pdf
Erstellt: 05.13 Revision: 05.15 Merkblatt Beschäftigte und Reisende Empfehlungen zur Vorbeugung und Notfallselbstbehandlung Dr. Gerhard Boecken Dr. Reinhard Krippner Malaria-Verbreitung und Infektionsrisiko Malaria-Risiko für Beschäftigte des Auswärtigen Amtes
Deep-Sea Research I 48 (2001) 405}437 Physical-biological coupling in the Algerian Basin (SW Mediterranean): In#uence of mesoscale instabilities on the biomass and production of phytoplankton and bacterioplankton XoseH Anxelu G. MoraHn �*, Isabelle Taupier-Letage�, Evaristo VaHzquez-DommHnguez , SimoHn Ruiz�, Laura Arin , Patrick Raimbault�, Marta Estrada Dept. Biologia Marina i Oceanograxa, Institut de Cie%ncies del Mar, CSIC, Pg. Joan de Borbo&, s/n, E-08039 Barcelona, Spain






